Psoriasis vulgaris is a chronic inflammatory skin disease, characterized by severe inflammation resulting in poor differentiation and hyperproliferation of keratinocytes [1]. Psoriasis is now considered as an independent risk factor of Cardiovascular Disease (CVD), as both psoriasis and atherosclerosis are T-helper Th1/Th17 mediated inflammatory diseases [2–4]. Recently, the concept of “psoriatic march” has been proposed as a cascade of events starting from inflammation of the skin, predisposing to systemic inflammation, increased oxidative stress, insulin resistance, endothelial dysfunction, and atherosclerosis and ultimately leading to CVD [5].
C-reactive protein (CRP) is an acute phase reactant, which is important for immune system activation during inflammation. Elevated level of hs-CRP is found in low grade systemic inflammation [6]. Its levels are genetically determined by 30-50% heritability [7]. CRP is considered as a marker of inflammation in several conditions including psoriasis and atherosclerosis [8]. Single Nucleotide Polymorphisms (SNP) of CRP such as +1846C/T rs1205 are associated with its serum level and co-morbidities of psoriasis [9,10]. Though increased CRP levels are seen in psoriasis, till date only one SNP association study was done in Taiwanese psoriatic subjects, which did not show any association [8,11]. Hence, the present study aimed to demonstrate the association of the variant rs1205 in the CRP gene involved in the pathogenesis of psoriasis with susceptibility to the disease and protein levels in South Indian Tamil patients with psoriasis.
Materials and Methods
This study included 300 patients of Tamil ethnicity (age range from 18 to 65) with psoriasis and 300 age, gender and ethnicity matched controls at Jawaharlal Institute of Postgraduate Medical Education and Research (JIPMER), Puducherry, India, from February 2014 to January 2016. It was approved by the JIPMER Institute Ethics Committee (Human Studies). The study was performed according to the World Medical Association Declaration of Helsinki ethical principles for medical research involving human subjects [12]. Written informed consent was obtained from all the participants of the study. Consecutive patients with psoriasis (age range from 18 to 65) constituted the case group. Pregnant women, individuals with other inflammatory and infectious disorders and patients on any systemic therapy in the last one month prior to recruitment, were excluded from the cases. International psoriasis council consensus classification of psoriasis was used for classification of psoriasis [13], and CASPAR criteria [14] was used for psoriatic arthritis. Disease severity was assessed by Psoriasis Area Severity Index (PASI) scoring [15]. A total of 300 age, gender and ethnicity-matched healthy individuals were recruited as healthy controls. Exclusion criteria for controls were individuals with inflammatory/infectious disorders, diabetes, hypertension, CVD, peripheral artery disease, family history of psoriasis or autoimmune disease.
Sample size was estimated at 5% level of significance with 80% power for the estimated disease prevalence of 0.8% [16] and minor allele frequency of 0.29 [17]. The sample size was calculated using CaTS Power Calculator Software (Power Calculator for Genetic Studies compiled and published by the Centre for Statistical Genetics, Michigan University, USA). The estimated sample size was 210 cases and 210 controls. The sample size was further modified to 300 subject in each group for enabling different trait sub group analysis.
Statistical analysis was performed by Graph Pad Instat 3 (Graph Pad Software Inc., San Diego, CA, USA). The observed allele frequencies in cases and controls were compared with expected frequencies to check for the Hardy–Weinberg Equilibrium (HWE) by Chi-square method. The frequency of genotypes and alleles were determined by the direct gene counting method. Within patient group, the frequency of genotypes and the alleles were correlated with clinical manifestations and disease activity. To examine the effect of the gene polymorphism on the protein concentrations, ANOVA was performed. Odds Ratio (OR) and Confidence Intervals (CIs) were calculated by Chi-square test. A two-sided p-value < 0.05 was considered as significant.
Genomic DNA extraction was performed by phenol-chloroform method [18]. Genotyping of CRP SNP rs 1205 was done by using TaqMan 5’allele discrimination assay. Allele specific fluorogenic probes (predesigned by Applied Biosystems Corporations, USA) amplified the target gene in a CFX96 TouchTM real time Polymerase Chain Reaction (PCR) detection system (Bio-Rad Laboratories, Inc., California, USA). The assay was performed according to the manufacturer’s protocol.
Homozygous major and heterozygous minor variant genotypes were determined based on fluorescent dyes (FAM and VIC). Thirty percent of the random samples were replicated for validation. Plasma levels of the high sensitivity C-reactive protein (hs-CRP) were estimated using commercially available ELISA kits (Calbiotech Inc, California, USA). The analytical range for the hs-CRP ELISA kit was 0.5-10 mg/L and the sensitivity was 0.005 mg/L. The intraassay variation was 5% and the interassay variation was 8%.
Results
We recruited 300 cases with psoriasis and 300 age, gender and ethnicity matched controls. Amongst cases, 214 were males and 86 females (mean age 43.56±12.10) and in controls, 210 males and 90 females (mean age 42.36±13.18) were recruited. Mean PASI score was 7.18 ± 6.69. Amongst cases, 230 had mild disease (PASI<10), and 70 had moderate-to-severe (PASI≥10) psoriasis. Mean age of onset was 37.31±12.98 years; 169 were early onset case (<40 years) and 131 were late onset case (≥40years). Mean disease duration was 73.87 ± 84.64 months. Thirty one patients had family history of psoriasis. Amongst cases, 231(77%) had chronic plaque psoriasis, 33(11%) psoriatic erythroderma, 2(0.67%) guttate psoriasis, 21(7%) palmoplantar psoriasis, 2 (0.67%) plantar psoriasis and 11 (3.67%) pustular psoriasis. One hundred and three (34.33%) patients had psoriatic arthritis and 132 (44%) had nail involvement.
The comparison of genotype and allele frequencies of rs1205 between patients with psoriasis and controls did not yield any significant difference, hence there is a lack of association of psoriasis risk and CRP SNP rs1205 [Table/Fig-1]. The genotypes of the cases and the controls were concordant with the HWE. Further we stratified the cases based on the clinical phenotype and observed lack of association of rs1205 with these clinical phenotypes [Table/Fig-2].
Genotype and allele frequencies of CRP [+1846 (C/T)] rs1205] in patients with psoriasis and healthy controls
| Gene & SNP | Genotype/allele | Psoriasis* (n=300) | Controls$ (n=300) | p-value | OR (95%CI) |
|---|
| CRP [+1846 (C/T)] | CC | 124 (41%) | 136 (45%) | | |
| CT | 136 (45%) | 123 (41%) | 0.31 | 1.21 (0.86-1.71) |
| TT | 40 (14%) | 41 (14%) | 0.89 | 1.07 (0.65-1.76) |
| C | 384 (64%) | 395 (66%) | 0.55 | 1.09 (0.85-1.37) |
| T | 216 (36%) | 205 (34%) | | |
* Psoriasis cases in Hardy Weinberg equilibrium (χ2=0.08, p=0.78); $Controls in HWE (χ2=2.35, p=0.13).
Influence of CRP (rs1205) on the phenotype of psoriasis.
| S. No. | Heading | CRP (rs1205) | No. of Positives (N) | No. of Negatives (N) | p-value* | OR | 95% CI |
|---|
| 1 | Females | CC vs CT+TT | 46 | 130 | 0.31 | 1.35 | 0.81 - 2.23 |
| CT | 36 | 100 | 0.37 | 1.32 | 0.77 - 2.26 |
| TT | 10 | 30 | 0.50 | 1.43 | 0.64 - 3.21 |
| CC | 40 | 84 | | | |
| 2 | Family History | CC vs CT+TT | 16 | 160 | 0.52 | 0.73 | 0.35 – 1.53 |
| CT | 13 | 123 | 0.65 | 0.77 | 0.35 -1.69 |
| TT | 3 | 37 | 0.60 | 0.59 | 0.16 – 2.15 |
| CC | 15 | 109 | | | |
| 3 | Early onset type | CC vs CT+TT | 104 | 72 | 0.30 | 1.31 | 0.83 – 2.08 |
| CT | 78 | 58 | 0.50 | 1.22 | 0.75 – 1.99 |
| TT | 26 | 14 | 0.23 | 1.69 | 0.80 – 3.53 |
| CC | 65 | 59 | | | |
| 4 | Psoriatic arthritis involvement | CC vs CT+TT | 56 | 120 | 0.33 | 0.76 | 0.47 – 1.24 |
| CT | 39 | 97 | 0.15 | 0.66 | 0.39 – 1.11 |
| TT | 17 | 23 | 0.74 | 1.21 | 0.59 -2.50 |
| CC | 47 | 77 | | | |
| 5 | Nail Involvement | CC vs CT+TT | 86 | 90 | 0.057 | 1.62 | 1.01 – 2.59 |
| CT | 66 | 70 | 0.083 | 1.60 | 0.97 – 2.63 |
| TT | 20 | 20 | 0.21 | 1.70 | 0.83 – 3.48 |
| CC | 46 | 78 | | | |
| 6 | Severity | CC vs CT+TT | 42 | 134 | 0.90 | 0.93 | 0.54 – 1.61 |
| CT | 31 | 105 | 0.97 | 0.99 | -0.55 – 1.77 |
| TT | 11 | 29 | 0.67 | 0.77 | 0.34 – 1.73 |
| CC | 28 | 96 | | | |
*Chi-square test.
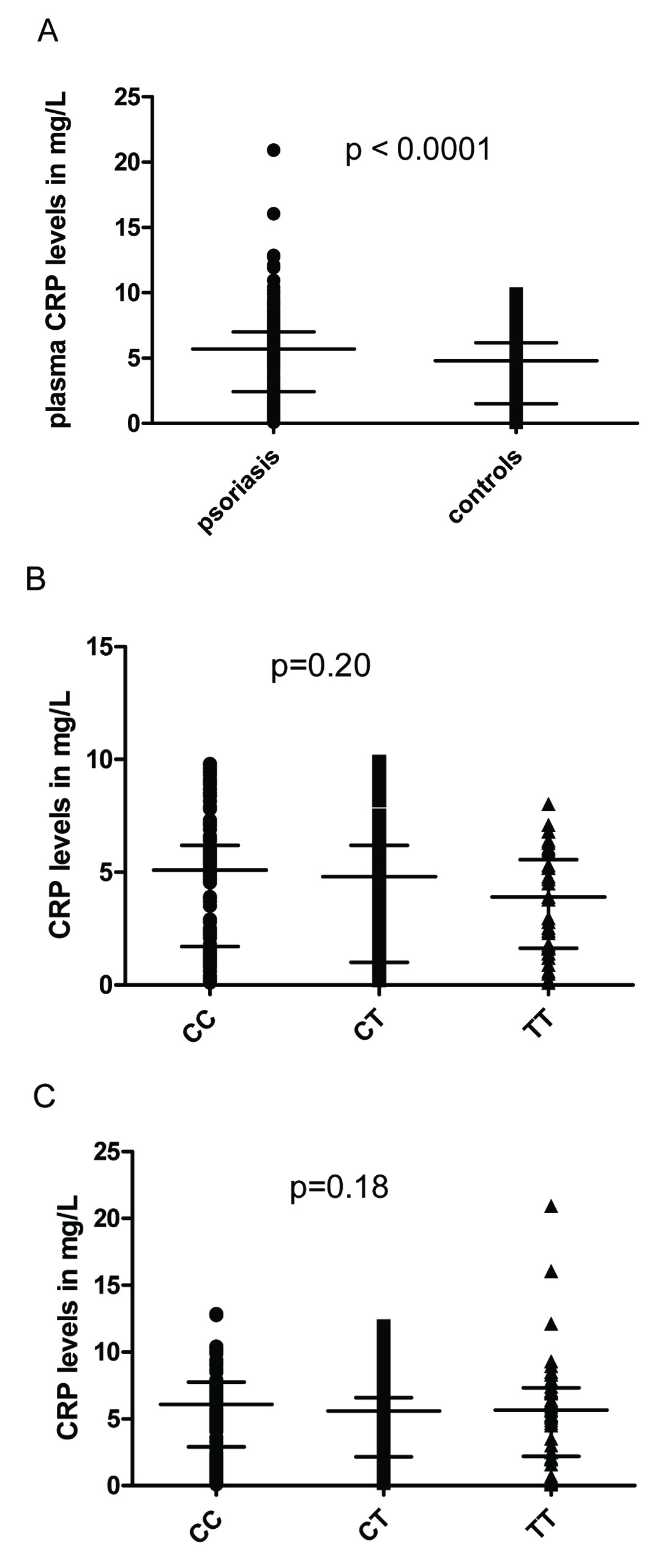
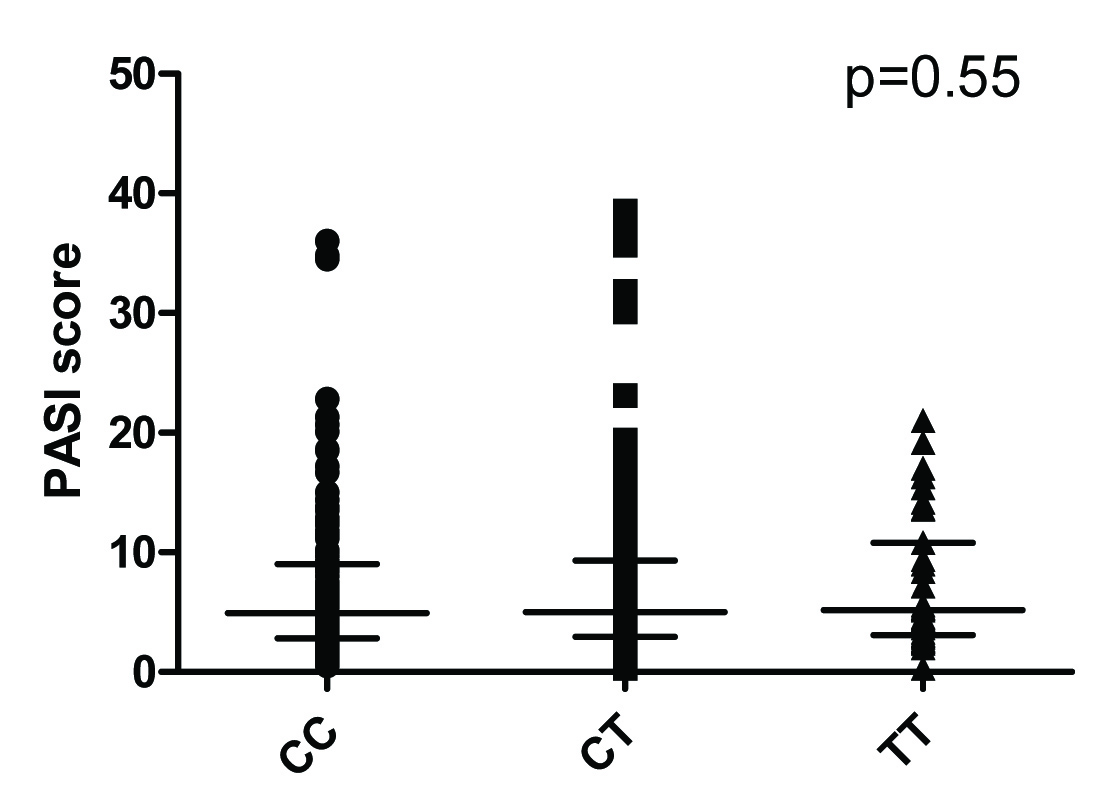
Plasma hs-CRP levels were significantly higher in cases than in controls [Table/Fig-3a]. Circulating hs-CRP levels correlated with disease severity (r=0.17, p=0.004). Further, CRP levels in both groups were stratified based on genotypes and analyzed for association with the CRP rs1205 genotype [Table/Fig-3b&c]. PASI score in cases were also stratified based on genotypes and analysed for association with CRP rs1205 [Table/Fig-4]. None of the genotypes were found significantly associated with PASI or with CRP levels in the study population.
Plasma hs-CRP levels in the study population.
(a) Plasma hs-CRP levels in patients with psoriasis vs. controls.
(b) Influence of rs1205 genotypes in CRP levels in controls.
(c) Influence of rs1205 genotypes in CRP levels in psoriasis.
Influence of genotypes on severity of psoriasis.
Discussion
In the present study, we investigated whether the genetic variant of CRP rs1205 was associated with susceptibility to psoriasis in the hitherto unexplored genetically distinct South Indian Tamil population. Our findings suggested that there is no association of CRP genetic variant rs1205 with psoriasis risk and CRP levels in South Indian Tamils, albeit hs-CRP levels were higher in cases and correlated positively with disease severity.
CRP gene is located on chromosome 1q23.2, in a conserved genetic region, which codes for proteins, with two exons separated by one intron coding GT repeat. Exon 1 codes for 18 amino acid leader peptide and the first two amino acids of the mature protein. The second exon encodes the remaining 204 amino acids, followed by a stop codon. CRP gene is highly polymorphic and all of the known CRP gene SNPs are in high linkage disequilibrium [19]. Certain genetic variants in the inflammatory biomarker genes such as CRP lead to greater relative production of these biomarkers in some individuals, rendering them more sensitive in response to the same inflammatory stimuli, than carriers of other variants. Many SNPs in the CRP gene were found associated with CRP levels independently, or in combination with another SNP in co-morbidities of psoriasis such as CVD in different races [9,10].
Circulating CRP present in the plasma comes from the liver, where its synthesis of CRP is mainly regulated by interleukin-6 (IL-6). IL-6 secretion is further up-regulated by pro-inflammatory cytokines such as IL-1 and tumour necrosis factor (TNF-α) [20]. CRP is also produced locally in atherosclerotic lesions by smooth muscle cells, lymphocytes and monocytes and considered as a robust marker of inflammation, future predictor of coronary heart disease and atherosclerotic events [10].
In the only published work involving CRP gene polymorphisms and psoriasis risk, Chang et al., observed that CRP SNPs were not associated with psoriasis susceptibility in a Chinese-Taiwanese cohort [11]. To the best of our knowledge, this work is a maiden venture in this area of research to investigate the association between CRP rs1205 and psoriasis in the ethnically distinct relatively unexplored South Indian Tamil population. Our study reports C as major allele for the CRP SNP rs 1205 in South Indian Tamils. We did not find significant association of the CRP SNP rs1205 with psoriasis and CRP levels in South Indian Tamils. CRP levels were higher in cases and showed significant correlation with disease severity, pointing towards a systemic inflammatory state in patients with psoriasis.
A 3’ Untranslated Region (UTR) SNP rs1205 has been studied in number of studies in different diseases [20–24]. Association of rs1205 with co-morbidities of psoriasis like myocardial infarction and stroke also varies in different populations [20–24]. Papaoikonomou et al., and co-workers reported that T allele is associated with vascular co-morbidities in diabetes [21]. In another study in Russian population, the SNP rs1205 was associated with coronary heart disease [22]. González-Giraldo et al., in a metanalysis of the association of CRP genetic variants with risk of ischemic stroke, reported that there was no significant association of CRP SNP rs1205 with ischemic stroke [23]. rs1205 genetic variant was associated with risk of myocardial infarction and the TT genotype was associated with lower levels of plasma CRP, compared with C allele carriers in a multicentric European study in patients with myocardial infarction [24].
Inconsistent results in inflammatory conditions in different populations may be due to different reasons. Highly polymorphic nature of the CRP gene and the fact that CRP SNPs are in high linkage disequilibrium may be one of the reasons [19]. CRP is regulated by many factors like IL-6, IL-1 and TNF-α [20]. Genetic variations in these regulatory molecules also affect the clinical phenotypes [22]. The differences in environmental exposure may also lead to variable risk due to systemic inflammatory response in psoriasis and its co-morbidities.
Limitation
A major limitation of this research work was that we did not have a replication cohort, which would have validated our results further. Secondly, had we analyzed multiple variants of the CRP gene, it would have bolstered our findings further, as the SNPs tend to act in conjunction in specific linkage blocks to exert their influence on the disease.
Conclusion
To conclude, we observed that plasma CRP levels are higher in patients with psoriasis and correlate with disease severity, whilst CRP rs1205 is not associated with susceptibility to psoriasis in our ethnic South Indian Tamil population. However, further studies are needed involving multiple genetic variants in the CRP gene to examine the impact of this inflammatory gene on the disease process in psoriasis.
* Psoriasis cases in Hardy Weinberg equilibrium (χ2=0.08, p=0.78); $Controls in HWE (χ2=2.35, p=0.13).
*Chi-square test.