The female vagina is a complex habitat for micro-organisms. Lactobacilli are the most dominant bacteria inhabiting this environment, as they play a major role in the maintenance of a healthy urogenital tract [1].
The microbial ecosystem in healthy “normal” vagina can be disrupted by drugs or by local devices with the concomitant decrease of the Lactobacilli community and overgrowth of anaerobic and facultative aerobic bacteria and yeasts [2].
Bacterial Vaginosis (BV) is one of the most common vaginal syndromes affecting women (pregnant or non- pregnant) in their reproductive age. It has been associated with a significant number of obstetric and gynaecologic complications.
Pregnant women with BV are more likely to have a preterm labour and delivery, preterm premature rupture of membranes, post- caesarean delivery wound infections, spontaneous abortion, chorioamnionitis, postpartum endometritis and subclinical pelvic inflammatory disease [3,4]. The mainstay treatment of BV consisted of metronidazole, clindamycin or tinidazole [5].
Many studies have considered probiotics as an adjunct to antimicrobial treatment [6–8]. Others proposed the use of probiotics in preventing the recurrence of BV after an initial treatment [8,9]. In addition, the prophylactic use of probiotics in healthy women with a history of recurrent BV has been confirmed [10].
Culture independent methods based on 16S rRNA-encoding gene sequence community profiling have been used previously to overcome culture-dependent limitations [11] and offer the potential to be rapid and reliable for characterizing the vaginal flora in clinical studies [12].
Our purpose was to assess a potential correlation between the taxonomy of the Lactic Acid Bacterial (LAB) species colonizing the vagina and a balanced healthy vaginal flora by comparing the vaginal LAB taxa found in healthy women and women with bacterial vaginosis by using Polymerase Chain Reaction (PCR) targeting universal and LAB specific 16S rRNA primer sets in conjunction with denaturing gradient gel electrophoresis (DGGE).
To our knowledge, this is the first report describing the vaginal lactic acid bacteria in Algerian pregnant women using this approach.
Materials and Methods
Swab Sampling, Nugent Score and pH Measurement
Vaginal samples from fifteen women were collected. Three swabs were collected from each subject after insertion of an un-lubricated speculum into the posterior fornix of the vaginal tract. Women were pregnant at various months of pregnancy and under no antimicrobial or prescribed therapy, aged between 19 and 35 years (mean 27.6 years; n=15). Swabs were vigorously agitated in 1mL sterile distilled water. One drop was deposited on microscope slide smears, Gram-stained, and scored by the Nugent method [13]. Nugent results were graded as 0–3 (Normal vaginal state; N), 4–6 (Intermediate-grade bacterial colonization; I), and 7–10 (Bacterial vaginosis; BV). The pH was determined using a pH indicator paper (Merck).
DNA Extraction and PCR Amplification
Microbial cell suspensions from vaginal swabs were pelleted by centrifugation (10,000 g, 5 min) previous to DNA extraction using ABIO pure Genomic DNA blood/cell culture (Alliance Bio Laborato-ries) according to manufacturer’s instructions. DNA was quantified using the NanoDrop microvolume sample retention system (Thermo Scientific NanoDrop Products). PCR reactions were carried out in a thermocycler (Thermal Cycler; Biometra) using the universal primers of bacteria 357F (5’-119 TACGGGAGGCAGCAG-3’) and 907R (5’-CCGTCAATTCMTTTGAGTTT-3’) as described by Muyzer [14] and the specific lactic acid bacteria primers Lac1 (5’-AGCAGTAGGGAATCTTCCA-3’), Lac2 (5’-ATTYCACCGCTACACATG-3’) and Lac3 (5’-AGCAGTAGGGAATCTTCGG-3’). A 44-bp GC-rich clamp sequence was added to the 5’ ends of primers 357FGC and Lac 2GC [15,16].
The PCR reaction was obtained by mixing 0.6 μL of each primer (25 mM), 10 μL buffer 5X with 20 mM MgCl2, 0.5 μL dNTPs (25 mM), 1 U/ μL of Taq polymerase (Thermo Scientific), 1μL of the genomic DNA, and ultrapure water (Millipore) to a final volume of 50μL. Amplified DNA was analysed after electrophoresis in 1.5% agarose gel followed by staining with ethidium bromide (5μg/ml) using Gel Compare (Version 4.1, Applied Maths, Kortrijk Belgium).
Taxa Identification using DGGE and 16S RNA Sequencing
PCR products generated with primers set N°1, 2 and 3 [Table/Fig-1] were separated by DGGE according to the specifications of Muyzer [15] with the following modifications: The concentrations of polyacrylamide, denaturant (7M urea, 40% formamide) and Tris-acetate buffer were 8%, 35–45% and, 50X respectively. A 20 μL of amplified PCR products mixed with 2X loading buffer were deposited into each well. Gel was run for 16 hour at 100V in TAE 1X buffer at 60°C in INGENYphorU system.
PCR conditions prior to DGGE community profiling.
| Cycle stage | Primer set N°1907R-357F GC | Primer set N°2Lac1-Lac2 GC | Primer set N°3Lac3-Lac2 GC |
|---|
| Initial denaturation | 94°C; 3 min | 94°C; 2 min | 94°C; 2 min |
| Cycles of denaturation | 10 cycles; 61°C; 1min25 cycles; 56°C; 1min | 35 cycles; 51°C; 1min | 35 cycles; 61°C; 1min |
| Elongation | 72°C; 1min | 72°C;1 min | 72°C; 1min |
| Final extension | 72°C; 7 min | 72°C; 7 min | 72°C; 7 min |
After electrophoresis, the gel was stained for 20 minutes in 5 μg/mL of ethidium bromide, washed with deionized water. Under UV light, DNA bands were cut from the gel, and placed in sterile 50 μL distillated sterile water overnight at 4°C.
A 5 μL of the solution containing fragments from the previously excised bands was used as template for PCR re-amplification as described above for DGGE except that primer sets lacking the GC clamps were used. The resulting PCR products were subsequently sequenced using the BigDye Terminator cycle sequencing kit V3.1 (Applied Biosystems) and an Applied Biosystems 3130 Capillary DNA Sequencer. Analysis of the partial 16S rRNA sequences was conducted using the GenBank DNA database NCBI and the BLAST algorithm [17].
Results of PCR and DGGE fragments in samples from women with Nugent ratings of N, I, or BV were presented as mean±SD.
Results
Nugent Score and pH Measurement of Vaginal Swabs
Among 15 pregnant women tested Nugent ratings for intermediate flora (8 cases), bacterial vaginosis (4 cases) and healthy flora (3 cases). Among the vaginal swabs of women with intermediate Nugent ratings, the pH was comprised between 4 and 5, while the pH of swabs of women with normal flora and bacterial vaginosis were of pH 4 and pH 5, respectively [Table/Fig-2].
Phenotypic characteristics of the vaginal flora of 15 pregnant women.
| Subject | Age (years) | Nugent score | Nugent rating | Vaginal pH |
|---|
| W01 | 24 | 6 | Intermediate flora | 4 |
| W02 | 26 | 3 | Normal flora | 4 |
| W04 | 21 | 5 | Intermediate flora | 5 |
| W05 | 35 | 7 | Bacterial vaginosis | 5 |
| W06 | 25 | 3 | Normal flora | 4 |
| W09 | 26 | 4 | Intermediate flora | 4 |
| W11 | 33 | 6 | Intermediate flora | 5 |
| W12 | 25 | 8 | Bacterial vaginosis | 5 |
| W13 | 27 | 3 | Normal flora | 4 |
| W14 | 22 | 4 | Intermediate flora | 5 |
| W15 | 19 | 5 | Intermediate flora | 4 |
| W16 | 32 | 6 | Intermediate flora | 5 |
| W17 | 27 | 5 | Intermediate flora | 4 |
| W18 | 24 | 7 | Bacterial vaginosis | 5 |
| W19 | 34 | 7 | Bacterial vaginosis | 5 |
The phenotypical characterization was carried out by determining the pH and the Nugent score for each swab.
Taxonomical Identification of Vaginal Swabs
DGGE analysis of PCR products using Lactobacillus-specific primers and universal primers showed that samples typically yielded 2.4 ± 0.95 dominant fragments per sample. It was possible to identify 12 bacterial species.
In women with Nugent scores rated normal flora, 2.66 ± 0.94 dominant DNA fragments were observed, and those fragments, when sequenced, were homologous to Lactobacillus iners, Lactobacillus delbrukii and Streptococcus agalatiae.
In women with Nugent scores rated intermediate flora, 2.12 ± 0.92 dominant DNA fragments were observed, and those fragments, when sequenced, were homologous to Aerococcus urinae, Enterococcus feacalis, Enterococcus hirae, Enterococcus spp., Lactobacillus acidophilus, Lactobacillus iners, Lactobacillusjensenii, Lactobacillus johnsonii, Lactobacillus crispatus, and Streptococcus anginosus with predominance of Enterococcusfeacalis (33.25%).
In women with Nugent scores rated bacterial vaginosis, 2.75 ± 0.82 dominant DNA fragments were observed, and those fragments, when sequenced, were homologous to Enterococcusfeacalis,Enterococcus feacium, Enterococcus hirae, Lactobacillusacidophilus, uncultured Lactobacillus spp. and Streptococcus anginosus, with predominance of Enterococcus feacalis (40%).
The homology between the sequence obtained from the DGGE bands and the closest species from the database are given in [Table/Fig-3].
(Suite): 16S rRNA sequence BLAST analysis corresponding to DGGE gel excised fragments.
| Subjects | Nugent rating | DGGE fragments† | Primer set | Band number | Most closely related bacterial sequence | GenBank accession no. | Identity, % |
|---|
| W01 | I | 3 | Lac3-Lac2 | 12 | Enterococcus faecalis | JUNL01000323 | 98% |
| Lac3-Lac2 | 13 | Enterococcus hirae | ASVZ01000003 | 97% |
| Lac1-Lac2 | 39 | Lactobacillus crispatus | AY335500 | 99% |
| W02 | N | 2 | Lac1-Lac2 | 38 | Lactobacillus iners | KU726667 | 99% |
| 357F-907R | 48 | Lactobacillus iners | ADHG01000003 | 99% |
| W04 | I | 2 | Lac3-Lac2 | 11 | Aerococcus urinae | NC_015278 | 98% |
| Lac1-Lac2 | 35 | Lactobacillus iners | ADHG01000003 | 98% |
| W05 | BV | 4 | Lac3-Lac2 | 22 | Streptococcus anginosus | AZMF01000010 | 97% |
| Lac3-Lac2 | 23 | Enterococcus faecalis | KX073796 | 99% |
| Lac3-Lac2 | 25 | Enterococcus faecalis | JUNL01000323 | 99% |
| 357F-907R | 44 | Uncultured Lactobacillus spp. | KM250388 | 99% |
| W06 | N | 2 | 357F-907R | 45 | Lactobacillus iners | ADHG01000003 | 99% |
| Lac3-Lac2 | 10 | Streptococcus anginosus | AZMF01000010 | 97% |
| W09 | I | 1 | Lac3-Lac2 | 20 | Enterococcus faecalis | JUNL01000323 | 99% |
| W11 | I | 2 | Lac1-Lac2 | 32 | Lactobacillus acidophilus | AYUA01000121 | 98% |
| Lac3-Lac2 | 19 | Enterococcus faecalis | KR137541 | 99% |
| W12 | BV | 2 | 357F-907R | 53 | Lactobacillus crispatus | KU991819 | 99% |
| Lac3-Lac2 | 9 | Streptococcus spp. | JMBI01000046 | 96% |
| W13 | N | 4 | Lac1-Lac2 | 29 | Lactobacillus delbrueckii subsp. delbrueckii | BALP01000125 | 97% |
| Lac1-Lac2 | 30 | Lactobacillus delbrueckii subsp.delbrueckii | BALP01000125 | 99% |
| Lac1-Lac2 | 31 | Lactobacillus delbrueckii subsp. bulgaricus | NC_014727 | 99% |
| Lac3-Lac2 | 18 | Streptococcus agalactiae | JMBI01000046 | 98% |
| W14 | I | 2 | Lac1-Lac2 | 28 | Lactobacillus jensenii | GG704690 | 97% |
| Lac3-Lac2 | 7 | Enterococcus spp. | KB029381 | 80% |
| W15 | I | 4 | Lac3-Lac2 | 8 | Enterococcus faecalis | KX073789 | 99% |
| Lac1-Lac2 | 26 | Lactobacillus iners | ADHG01000003 | 100% |
| Lac1-Lac2 | 27 | Lactobacillus iners | KU726667 | 100% |
| Lac3-Lac2 | 5 | Streptococcus anginosus | KU726683 | 99% |
| W16 | I | 1 | Lac3-Lac2 | 17 | Enterococcus faecalis | JUNL01000323 | 99% |
| W17 | I | 2 | 357F-907R | 55 | Lactobacillus johnsonii | KU991818 | 99% |
| Lac3-Lac2 | 14 | Enterococcus spp. | KX079817 | 99% |
| W18 | BV | 3 | Lac3-Lac2 | 15 | Enterococcus faecium | KB946737 | 98% |
| Lac3-Lac2 | 16 | Enterococcus faecalis | JUNL01000323 | 99% |
| Lac3-Lac2 | 1 | Enterococcus faecalis | KK640484 | 99% |
| W19 | BV | 2 | Lac3-Lac2 | 2 | Enterococcus hirae | ASVZ01000003 | 97% |
| Lac3-Lac2 | 3 | Uncultured Lactobacillus spp. | KM250388 | 99% |
BV, bacterial vaginosis; I, Intermediate grade bacterial colonization; N, normal flora; DGGE, denaturing gradient gel electrophoresis.
† Approximate number of dominant fragments observed.
Several species of Lactobacillus were detected: Lactobacillus iners and Lactobacillus delbrueckii in the case of normal flora, Lactobacillus jensenii, Lactobacillus johnsonii, Lactobacilluscrispatus, Lactobacillus acidophilus and Lactobacillus iners in the case of intermediate flora and Lactobacillusacidophilus, uncultured Lactobacillus spp. in the case of bacterial vaginosis [Table/Fig-4].
Occurrence of Gram positive bacteria in the vagina of Algerian pregnant women.
The vaginal bacterial flora was classified according to the Nugent scores. Two levels of taxonomical classification are shown: at the level of the genus and of the species.
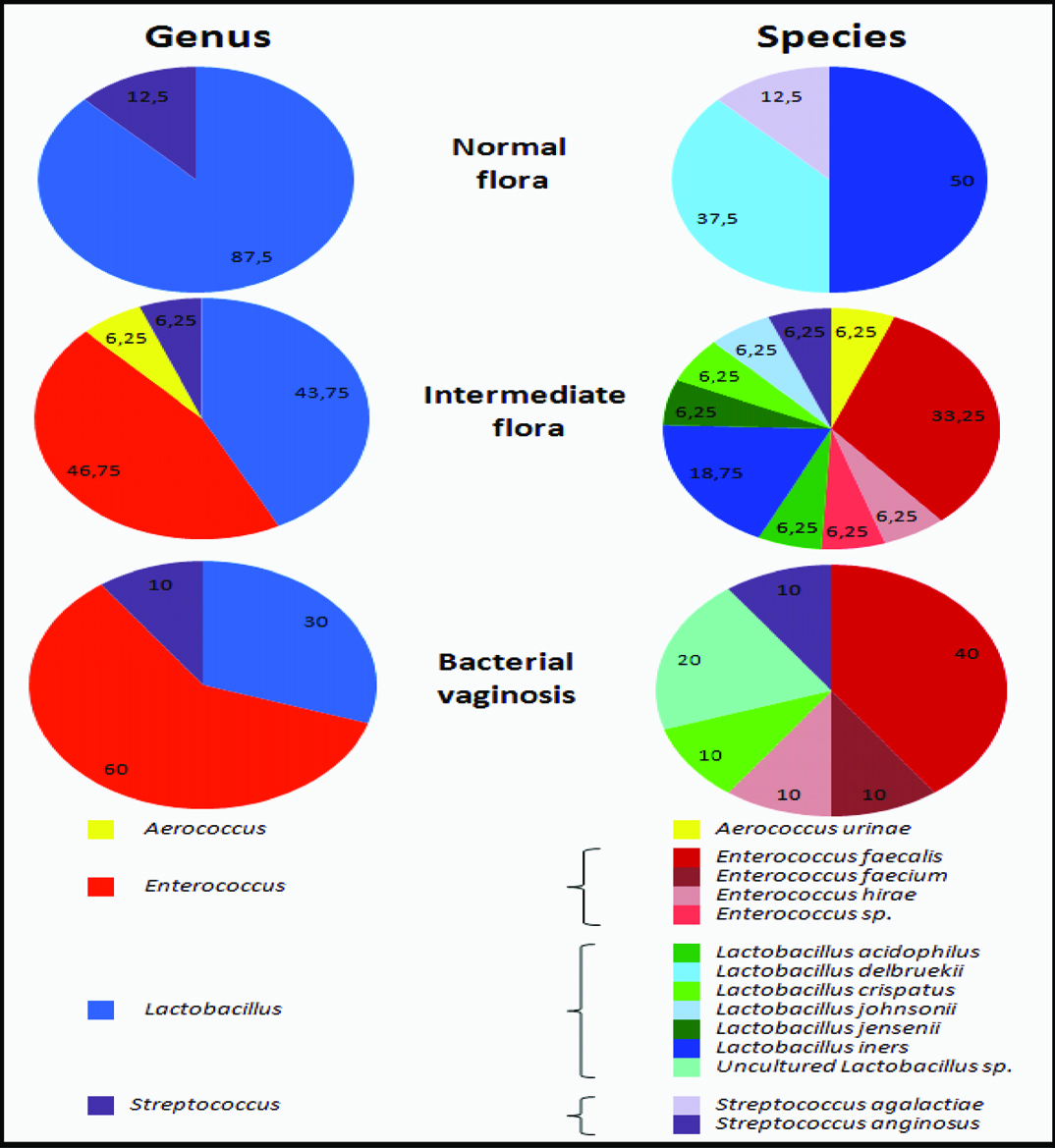
It was observed that the loss in detection of a Lactobacillus species by DGGE and sequencing analysis correlated with a higher Nugent score [Table/Fig-4]. In addition, all the women samples were colonized by a single species (or group of closely related strains) of Lactobacillus.
Our data suggest the absence of Enterococcus species in women belonging to normal flora, in contrast with others women belonging to intermediate flora or to bacterial vaginosis which are predominated by Enterococcus feacalis. So, this species can be considered as an indicator of imbalance of the vaginal ecosystem.
Discussion
The maintenance of a low pH in the vagina through the microbial production of lactic acid is known to be an important defense against infectious disease in reproductive age women [18]. In our study, all communities (women with normal flora, intermediate flora or bacterial vaginosis) contained members that have been assigned to genera known to produce lactic acid (including Lactobacillus, Streptococcus, Aerococcus and Enterococcus), but the correlation between this bacteria and the maintenance of a low pH has not been well defined.
Communities belonging to normal flora have the lowest median pH (4.0), whereas communities belonging to bacterial vaginosis have the highest median pH (5.0). Clearly, the acidification of vaginal ecosystem is negatively correlated with the incidence of bacterial vaginosis among women. These results are in agreement with those of Zhou et al., [19].
The findings of our study indicate that the structure of vaginal lactic acid bacteria varied between the three categories. The vaginal flora of each women were colonized by a single species of Lactobacillus, which is consistent with the findings of other studies [11,20,21].
In particular, L. delbrueckii detection in vaginal swabs of healthy women has been previously recorded but as a minor vaginal bacterial species [22] and not as a major flora as observed here (W13). We observed that vaginal swabs of healthy pregnant women were mostly colonized by L. iners (W02 and W06). The occurrence of L. iners as a member of vaginal communities in healthy women has also been demonstrated by others [19,23–26]. First described in 1999 [27], L. iners does not grow on Man Rogosa and Sharpe media (MRS agar). [14,19] which highlights the usefulness of culture-independent approaches for vaginal flora inventory.
Lactobacillus jensenii, Lactobacillus johnsonii, Lactobacilluscrispatus and Lactobacillus acidophilus were recovered from Algerian pregnant women with intermediate flora, in contrast with other studies that have frequently identified these species in vagina of women with normal flora [3,18,25,28–31].
Women belonging to intermediate flora were also characterized by a predominance of Enterococcus feacalis. In addition, Enterococcus hirae, Aerococcus urinae and Streptococcus anginosus have been identified in this category of women. These results are in correlation with other studies which confirmed the vaginal colonization of these women (intermediate flora) by aerobic enteric commensals [32,33].
In contrast with a general opinion that species of Lactobacillusacidophilus constitute most of the healthy vaginal Lactobacillus flora [25,28], our results recovered the absence of this species in a healthy women and its presence in 6.25 % of women with intermediate flora. These results suggests that this species is not indicative of a normal microflora, but may point to some intermediate condition.
According to research in PubMed, our study is the first to describe the diversity of lactic acid bacteria associated to bacterial vaginosis in contrast with the entire previous studies which describe only the pathogenic bacteria [34–36], or describe only Lactobacillus species associated to this category (women with Bacterial vaginosis) [37,38]. In addition, as per our knowledge our study is the first to describe the vaginal flora of Algerian women using culture independent method.
Our results need to be confirmed in a larger cohort preferably using a study design combined with data on subjects’ behaviors to avoid the limitations of the present study and properly reflect the diversity of vaginal flora in female population of Algeria.
Conclusion
Our data suggest that a predominance of Lactobacillus in the vaginal bacterial flora is negatively correlated with the incidence of bacterial vaginosis in 15 pregnant women of Algeria. In addition, Enterococcus genus is negatively correlated with the incidence of normal Nugent rating among the tested women. According to our results, L. iners and L. delbruekii can be considered as critical for defense of the vagina. In addition, Enterococcus feacalis can be defined as an indicator of the imbalance of vaginal ecosystem.
Compared to culture-dependent methods, PCR- DGGE technique able a better estimation of species composition of the vaginal community, which is a prerequisite for clinical studies of the vaginal tract. Future development of effective probiotics must take into account the results of this study which gives an idea on the specificity of the vaginal lactic acid bacterial flora of Algerian woman where local perineal hygiene is different from the western world.