Introduction
Cell culture technique was first developed in the early 20th century as a method of studying animal cell behavior in vitro [1]. The principle of cell culture was established when Roux, an embryologist used warm saline to maintain chicken embryo for several days, thereby, coming up with tissue culture principle [2]. Cell culture has therefore, been defined as the is removal of animal cells and its propagation and cultivation in vitro in an artificial environment that is suitable for its growth [3,4]. This usually begins with a primary culture aiming at achieving confluence, that is formation of monolayer of cell in a culture flask supplemented the required nutrients and growth factors. With achievement of confluence, the cells are then passaged or sub cultured from the primary to secondary and subsequence to tertiary, until a continuous cell line is established. The isolation of virus in a cell culture is labour-intensive, and consumes time [5,6]. Many clinically important viruses are still either difficult to grow or don’t grow at all in tissues culture while others may require a sophisticated culture system which may either not be suitable for diagnostic laboratory use or not available at all. These might reduce the impact of tissue culture in clinical diagnosis, thereby making it less attractive in diagnosing human diseases [5,7] while, some scientist found tissue culture as a relatively unbiased, whose limitation is only by the ability of the virus to grow on the selected cell lines [8,9]. However, Vero E6 cells were considered as the most permissive of all cell lines by providing a versatile medium for recovery of unknown pathogens, together with Electron Microscopy (EM) to the detection and classification of unknown agent [10,11].
Observation of cell culture via EM can provide early clues on aetiologic agent and subsequently guide laboratory and epidemiologic investigations. This is of clinical important once most specially, during diseases outbreak since knowing the aetiologic agent will assist public health officials to institute a timely response and prevent or limit further spread of the causative agent [12,13]. Therefore, the use of classical techniques of viral isolation in tissue culture and examination under EM is said to be critical in detection of viruses that were previously unrecognized as such. Contrary to the earlier view, cell culture is a fundamental technique that can be accomplished in hospital diagnostics and microbiology laboratories if infectious viral agent is suspected. This technique was used in discovering Ebola virus in a suspected yellow fever patient and vice versa in several studies [14–17].
Recent advances in metagenomics with deep sequencing techniques have made it possible to analyse the genome of microorganism without isolating the virus via cell culture. This is done via high-throughput sequencing using random amplified DNA product and comparison of sequences with available extensive bank of sequences for the final identification of the detected agent. This is possible because random primers can specifically amplify the template for sequencing without having a prior knowledge of the suspected agent [18–20]. This technique is readily advancing in the aspect of pathogen discovery. It has been used forever to discover viruses such as Lioviu virus [21], Schmallenberg virus [22] and Bas –Congo virus [23]. In the cases of severely ill patients or infectious diseases outbreak, it is important to identify the causative agent of infection. As such this review is aimed at describing some of the events in which viruses are isolated for identifying the causative agent and recognition of emerging diseases, by additional laboratory diagnosis assay such as Electron Microscope (EM), serological and molecular techniques.
Inoculation of clinical specimens from a patient on to the culture cells enables biological amplification of the virus to the level at which it can be detected or viewed under EM and further confirmed by other techniques such as serology, immunohistochemistry as well as fluorescence antibody assays and molecular methods leading to further characterization of the species and strain of the virus [24–26]. Therefore, culture based system for viral isolation have been the “gold standard” for the diagnosis of viral infections in clinical virology and have served the laboratory well for decades [27]. However, the use and relative importance of virus culture has been on the decline due to development of rapid and accurate molecular techniques [28–30]. Therefore, the aim of this review is to critically summarize the views of researchers on the role of cell culture technology in diagnosis of human diseases.
Methodology
Searches for peer-reviewed journal articles were conducted using the University Putra Malaysia online subscribed databases in the area of Health Sciences and Medicine via databases such as; Medline, SCOPUS and Google Scholar search engine. All searches were limited to publication from 2000 to 2015 except were necessary an older publication might be consider. All publications were in English and duplicates were removed. The final articles searched were those published till 31st May 2015. The online database serch resulted in 2473 articles which were screened base on the title and abstrcts relevance, exluding conference abstract, comments and short communications retaining 260 for full text review studies
Cell culture and electron microscopy in diagnosis
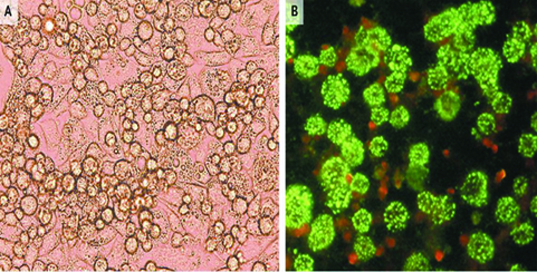
Electron Microscopy (EM) and cell culture isolation are instrumental in finding the causative agent in an unusual clinical manifestation. One of the studies reports the isolation of Bunya virus in patient with history of tick bites [31]. Initially, there was a suspicion of an Ehrlichiasp, therefore, leukocytes from the suspected patient were inoculated into DH82 cell (canine monocytes cell line) and it showed some cytological changes [Table/Fig-1]. The cells were then processed for examination with EM after which a Bunya virus was observed rather than the bacteria that was suspected. In cells infected with Bunya virus, the virus particles are found as a bud in vesicles and extracellular [Table/Fig-2]. The virus envelope is spherical with some projection on the surface of the virus particles and the virus has granular core.
a) Visible virus induced cytopathic effect in DH82 cells 8 days post infection with Bunya virus. The infected cells show visible granular particles and differentiated into macrophages with elongated pseudopodia. b) Bunya virus grown in vero cells, detected on immunofluorescenceassay [31].
a) Negative stain Bunya virus purified from infected Vero cells. b) Transmission electron microscopy of virus infected cells (DH82) shown by black arrows [31].
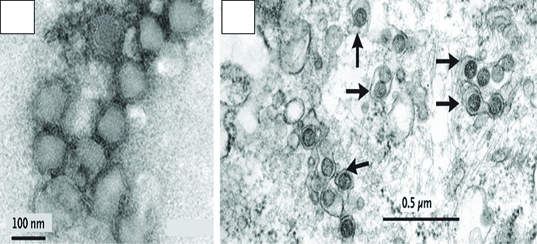
Cell Culture and RT– PCR
Cell culture and real time reverse transcription polymerase chain reaction (qRT-PCR) have been broadly used in clinical settings for identifying influenza viruses [32,33]. Although, time consuming and labor intensive and required high skill personnel with specific laboratory equipment and condition, making it not suitable for primary health care settings and low income countries. Nevertheless, cell culture is still important in confirming causative agent of infection in an outbreak. The currently reported H7 N9 cases of influenza infection were confirmed by cell culture and RT-PCR [34].
Cell Culture - Metabolomics
Cell cultures metabolomics can be used for identifying biomarkers of a pathological condition as well as metabolic path ways that produce such biomarkers. Metabolites play an important role in cancer diagnosis, recurrences and prognosis by identifying novel cancer biomarkers. A slight change in metabolism can be detected in products of cellular process leading to development of prognostic models that will be useful for early detection of cancer. Several studies examined the ability of human cancer cells to secrete volatile organic compounds [35,36], some of which were able to detect acetaldehyde release from lung cancer cell lines CALU-1 and SK-MES [37,38].
Rapid Detection Cell Culture
With the existence of commercially produced, cultivated cell lines which are used for rapid detection of a variety of viruses like R-Mix (Diagnostic hybrid, Inc) which is a mixture of monolayers of cells which are selected based on their capability of isolating different viruses causing respiratory tract infection. R-Mix contains tissue from the lungs (MV1LU) and A549 cells as fresh cells readily for use, or frozen cell suspension that can be aliquoted by the labouratory or as frozen monolayers in a shell vials ready to use. R mixed has therefore been reported to offer a fast and time sensitive technique of identifying viruses that are commonly involved in causing respiratory infection with no specialized skills required [39, 40].
Transgenic Cell Lines and Viral Detection
Transgenic technology in cell culture involves incorporation of stable genomic materials into the cell so that once a particular virus enters the cell, it triggeres, production of virus specific enzymes that is easily measurable [41,42]. The genetic materials can be of viral, bacterial or cellular origin and referred to as virus inducible reporter gene segment [43,44]. In diagnostic laboratory transgenic cells can only be useful if they have the desirable promoter which is quite in cells that are infected but significantly up regulated by means of viral trans-activator protein in a manner that is specific, but not allowing heterologous viral transactivation protein to stimulate the promoter. For a transgenic system to work, the virus to be detected must be able to adhere to the cell wall and prime its replication cycle without reaching the finishing point but adequate to activate the gene through the promoter. This makes the use of genetically improved cell line in improving growth of viruses possible, thereby, facilitating the detection of cells infected with viruses thus, providing the detection system that is specific, sensitive and very simple to perform [45,46]. This technology was applied successfully in identifying the polio virus via the use of transforming cells susceptible HeLa cells [47]. However, its identification is by staining with monoclonal antibodies and it can be detected within 16 to 24 hours inoculation [48,49]. Conversely, a more rapid transgenic system capable of easily detecting HSV within 24 was developed in such a way that it does not require medical expertise or expensive monoclonal antibodies. It involves the use of UL39 derived HSV promoter which codes for large ribo nucleotide reeducates sub unit [50,51].
Expression of recombinant protein for detection of influenza virus antibody
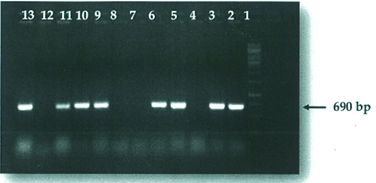
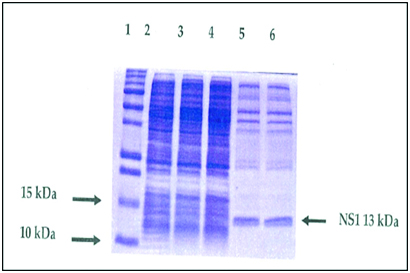
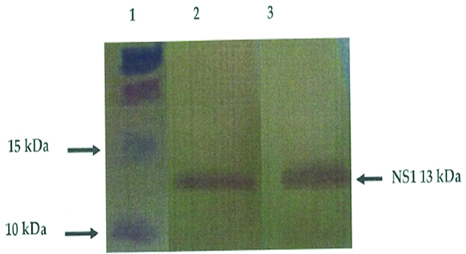
Recombinant protein technology is important in meeting the demand for easy to use, fast and reliable test in diagnostic laboratory and has been useful for serological survey of infection [52]. Recombinant protein can be expressed and used for the detection of influenza virus antibody. For instance, NSI gene was purified and cloned successfully into a vector [pCR2.1 TOPO TA cloning (3.9 kb], and then transformed into a competent cells (TOPOIO F’ E. coli strain) spread on LB agar and incubated at 37°c over night. The positive colonies containing the NSI gene was screened using PCR. The results showed an expected band of 690bp on agarose gel [Table/Fig- 3] [53]. It was sequenced and confirmed to be in frame with the N-terminal, together with proper orientation. The recombinant plasmids were then transformed into the host cell strain B12 (DE3) pLysS for expression. Transformation process was achieved using heat shock method. The expression of the expressed protein was analysed by SDS–PAGE [Table/Fig-4] which was further confirmed by western–blotting with the expected 13 KDa protein being immunocreatives to by poly clonal anti-NS antibody [Table/Fig-5] [54]. This confirmed that the antigen could be used to detect specific antibodies against influenza viruses using ELISA, which has a considerable advantage over other techniques for detecting specific antibodies.
Result of successful positive recombinant colonies ligation. Lane 1: DNA ladder 1kb; Lane 2-13: positive colonies with NS1; Lane 4, 7, 8 and 12: negative colonies with no NS1 inserted [53].
SDS-PAGE profile of un-purified expressed NS1 (13KDa) protein and un-induced plasmid. Lane 1: protein ladder; Lane 2 - 4: un-induced recombinant plasmid at zero hour; Lane 5 and 6: un-purified recombinant NS1 (13KDa) protein [54].
Result of western blot analysis of the purified proteins from E. coli containing pRSET B/NS1. Lane 1: protein ladder; Lane 2 and 3: purified recombinant NS1 protein [54].
Issues arising from this review
Standardization
Unlike molecular techniques, cell culture results can considerably differ, depending on collection, transport as well as handling of the specimens to maintain viral viability and healthy inoculated cells [55]. Researchers argued for and against the importance of cell culture in clinical laboratories. While some believed that there will be situations that will warrant the use of tissue culture in diagnostic virology laboratory, others think that it may be true to some level but not at the point of care, therefore, changing the significance of cell culture in diagnostics [56,57]. However, the molecular quantitative assay is still highly variable as such required standardization [58,59]. Viral isolation could be done when needed for a definite purpose by selected local region as well as national laboratories that have the required expertise and maintain cell culture system [60].
Time Consuming
Cell culture is rapidly losing its place and its relative significance in the diagnosis of human diseases in this era of wanting an immediate and accurate clinical diagnostic needed for early and effective intervention. On the other hand, molecular techniques provide a timeless and accurate diagnostic method. Therefore, molecular techniques are becoming the new “gold standard” and rapidly displacing the traditional cell culture based, early and accurate diagnostic methods that have significant impact on patient care in limiting the extend of diseases via timely treatment, thereby, reducing unnecessary hospitalization, antimicrobial use and their associated cost [61].
Labour Intensive
Cell culture required expertises and trained technologists as well as sophisticated equipments. It is, therefore, important to use the available technology based on a particular situation which will yield more useful result. With the use of transgenic technology pathogen identification was found to be successful, however, they are relatively labour intensive and required expertise. With the development of rapid cell culture technique that uses fluorescence staining, in which color change is used to identify pathogens, the need for intensive labour was reduced because the technologist don’t have to be skilled in maintaining CPE in cells [62]. Therefore, laboratories should evaluate the required resources, facilities, level of training and expertise required.
Sensitivity
It is clear from this review that some researchers are of the view that cell culture is less sensitive than the molecular method like PCR, with a large restriction in the spectrum of viruses. Thereby, making cell culture less useful for the “non culturable“viruses, limiting its sensitivity for its use in diagnosis [63–65]. On the other hand some concerns over molecular methods like PCR are that of false negatives due to PCR inhibitors and genetic diversity of the viruses as well as false positive as a result of contamination latent infection and viral co-infection. It is, therefore, important to use cell culture in monitoring and evaluating the sensitivity and specificity of molecular methods.
General Overview
The authors for and against the use of cell culture in the diagnosis of human diseases did their research well and their results were well presented except for the discussion of the results which show clearly the myopic nature by which those against and for cell culture look at it. Therefore, my criticism is based on the following points: It is important to note that, the mere detection of pathogen using molecular or next generation sequencing technique is just the first step; there is a need to determine whether or not the identified pathogen is associated with diseases that can only be achieved via cell culture; It is clear from this review that there is no single optimal approach for viral detection in all clinical circumstances. It is, therefore, important to combine both, cell culture and molecular techniques, in optimizing diagnosis of viral infection to achieve a cost effective, labour saving and medically important viral testing.
With the emergence and re-emergence of new strains of viruses that are not detectable by the currently available molecular methods, it is important to emphasize on cell culture as the gold standard in disease discovery and causation.
Conclusion and Recomendations
In conclusion, cell culture is an indispensable tool in modern day medicine and its applications are innumerable in diagnosis of human infection. Cell culture methods are unbiased to some extent and only limited by the ability of the virus to grow in a particular cell line. However, this has been overcome with the advent of transgenic cell culture technology. We therefore, recomend: Cell culture should be used in monitoring specificity and sensitivity of rapid tests based upon antigen assays annually and the clinicians should be notified of the results; Cell culture should also be encourage for negative rapid test result obtained from patient features infection during high prevalence or outbreak as well as for positive result during low prevalence: Cell culture can also be used in combination with PCR serological test, histopathology and immune histochemistry for diagnosis of unknown virus. They are also used in establishing a rapid test for newly discovered pathogens.
[1]. Thorpe TA, History of plant tissue cultureMolecular Biotechnology 2007 37(2):169-80. [Google Scholar]
[2]. Fung S, Wong F, Hussain M, Lok A, Sustained response after a 2-year course of lamivudine treatment of hepatitis B e antigen-negative chronic hepatitis BJournal of Viral Hepatitis 2004 11(5):432-38. [Google Scholar]
[3]. Ganem D, Schneider RJ, Hepadnaviridae: the viruses and their replicationFields Virology 2001 2:2923-69. [Google Scholar]
[4]. Willmer EN, Cells and tissues in culture: methodsBiology and Physiology: Elsevier 2013 [Google Scholar]
[5]. Leland DS, Ginocchio CC, Role of cell culture for virus detection in the age of technologyClinical Microbiology Reviews 2007 20(1):49-78. [Google Scholar]
[6]. Oberste MS, Nix WA, Maher K, Pallansch MA, Improved molecular identification of enteroviruses by RT-PCR and amplicon sequencingJournal of Clinical Virology 2003 26(3):375-77. [Google Scholar]
[7]. Griffith LG, Naughton G, Tissue engineering—current challenges and expanding opportunitiesScience 2002 295(5557):1009-14. [Google Scholar]
[8]. Khademhosseini A, Vacanti JP, Langer R, Progress in tissue engineeringScientific American 2009 300(5):64-71. [Google Scholar]
[9]. Ransohoff RM, Perry VH, Microglial physiology: unique stimuli, specialized responsesAnnual Review of Immunology 2009 27:119-45. [Google Scholar]
[10]. Abiko C, Mizuta K, Itagaki T, Katsushima N, Ito S, Matsuzaki Y, Outbreak of human metapneumovirus detected by use of the Vero E6 cell line in isolates collected in Yamagata, Japan, in 2004 and 2005Journal of Clinical Microbiology 2007 45(6):1912-19. [Google Scholar]
[11]. Weidmann M, Sanchez-Seco MP, Sall AA, Ly PO, Thiongane Y, Lô MM, Rapid detection of important human pathogenic PhlebovirusesJournal of Clinical Virology 2008 41(2):138-42. [Google Scholar]
[12]. Goldsmith CS, Ksiazek TG, Rollin PE, Comer JA, Nicholson WL, Peret TC, Cell culture and electron microscopy for identifying viruses in diseases of unknown causeEmerging Infectious Diseases 2013 19(6):864 [Google Scholar]
[13]. Alberts B, Bray D, Hopkin K, Johnson A, Lewis J, Raff M, Essential cell biology: Garland Science 2013 [Google Scholar]
[14]. Kann M, Bischof A, Gerlich WH, Invitro model for the nuclear transport of the hepadnavirus genomeJournal of Virology 1997 71(2):1310-16. [Google Scholar]
[15]. McMullan LK, Frace M, Sammons SA, Shoemaker T, Balinandi S, Wamala JF, Using next generation sequencing to identify yellow fever virus in UgandaVirology 2012 422(1):1-5. [Google Scholar]
[16]. Muyembe-Tamfum J-J, Mulangu S, Masumu J, Kayembe J, Kemp A, Paweska JT, Ebola virus outbreaks in Africa: past and presentOnderstepoort Journal of Veterinary Research 2012 79(2):06-13. [Google Scholar]
[17]. Kuno G, 23 Yellow Fever VirusManual of Security Sensitive Microbes and Toxins 2014 :265 [Google Scholar]
[18]. Mokili JL, Rohwer F, Dutilh BE, Metagenomics and future perspectives in virus discoveryCurrent opinion in virology 2012 2(1):63-77. [Google Scholar]
[19]. Willner D, Hugenholtz P, From deep sequencing to viral tagging: recent advances in viral metagenomicsBioessays 2013 35(5):436-42. [Google Scholar]
[20]. Lagier J-C, Edouard S, Pagnier I, Mediannikov O, Drancourt M, Raoult D, Current and past strategies for bacterial culture in clinical microbiologyClinical microbiology reviews 2015 28(1):208-36. [Google Scholar]
[21]. Mehedi M, Ebola virus RNA editing: Characterization of the mechanism and gene products 2011 [Google Scholar]
[22]. Liew FY, Pitman NI, McInnes IB, Disease-associated functions of IL-33: the new kid in the IL-1 familyNature Reviews Immunology 2010 10(2):103-10. [Google Scholar]
[23]. Deshmane SL, Kremlev S, Amini S, Sawaya BE, Monocyte chemoattractant protein-1 (MCP-1): an overviewJournal of Interferon & Cytokine Research 2009 29(6):313-26. [Google Scholar]
[24]. Gardner PS, McQuillin J, Rapid virus diagnosis: Application of immunofluorescence: Butterworth-Heinemann 2014 [Google Scholar]
[25]. Straus D, Rapid and sensitive detection of cells and virusesGoogle Patents 2011 [Google Scholar]
[26]. Kango N, Textbook of Microbiology: IK International Pvt Ltd 2010 [Google Scholar]
[27]. De Serres G, Skowronski D, Wu X, Ambrose C, The test-negative design: validity, accuracy and precision of vaccine efficacy estimates compared to the gold standard of randomised placebo-controlled clinical trialsEuro Surveill 2013 18(37) [Google Scholar]
[28]. Emerson SU, Purcell RH, Hepatitis E virusReviews in Medical Virology 2003 13(3):145-54. [Google Scholar]
[29]. Yang S, Rothman RE, PCR-based diagnostics for infectious diseases: uses, limitations, and future applications in acute-care settingsThe Lancet infectious diseases 2004 4(6):337-48. [Google Scholar]
[30]. Niesters HG, Clinical virology in real timeJournal of Clinical Virology 2002 25:3-12. [Google Scholar]
[31]. Yu X-J, Liang M-F, Zhang S-Y, Liu Y, Li J-D, Sun Y-L, Fever with thrombocytopenia associated with a novel bunyavirus in ChinaNew England Journal of Medicine 2011 364(16):1523-32. [Google Scholar]
[32]. Templeton KE, Scheltinga SA, Beersma MF, Kroes AC, Claas EC, Rapid and sensitive method using multiplex real-time PCR for diagnosis of infections by influenza A and influenza B viruses, respiratory syncytial virus, and parainfluenza viruses 1, 2, 3, and 4Journal of Clinical Microbiology 2004 42(4):1564-69. [Google Scholar]
[33]. Rahman M, Vandermause MF, Kieke BA, Belongia EA, Performance of Binax NOW Flu A and B and direct fluorescent assay in comparison with a composite of viral culture or reverse transcription polymerase chain reaction for detection of influenza infection during the 2006 to 2007 seasonDiagnostic Microbiology and Infectious Disease 2008 62(2):162-66. [Google Scholar]
[34]. Xia J, Liu L, Wang L, Zhang Y, Zeng H, Liu P, Experimental infection of pregnant rabbits with hepatitis E virus demonstrating high mortality and vertical transmissionJournal of Viral Hepatitis 2015 [Google Scholar]
[35]. Zhao W-D, Chen J, LIU F-G, Wang M, LI J-M, Poster Abstracts–LiverChinese Journal of Digestive Diseases 2005 6:A31-A51. [Google Scholar]
[36]. Beebe K, Employing metabolomics in cell culture and bioprocessing to gain greater predictability, control and qualityAnnual Meeting and Exhibition 2014 (July 20-24, 2014) 2014Simb [Google Scholar]
[37]. Smith D, Wang T, Sulé-Suso J, Španel P, Haj AE, Quantification of acetaldehyde released by lung cancer cells invitro using selected ion flow tube mass spectrometryRapid communications in mass spectrometry 2003 17(8):845-50. [Google Scholar]
[38]. Kalluri U, Naiker M, Myers M, Cell culture metabolomics in the diagnosis of lung cancer—the influence of cell culture conditionsJournal of breath research 2014 8(2):027109 [Google Scholar]
[39]. Tatematsu K, Tanaka Y, Kurbanov F, Sugauchi F, Mano S, Maeshiro T, A genetic variant of hepatitis B virus divergent from known human and ape genotypes isolated from a Japanese patient and provisionally assigned to new genotype JJournal of Virology 2009 83(20):10538-47. [Google Scholar]
[40]. Huang YT, Transgenic mink lung cells (which express human furin) which show increased sensitivity to infection or are capable of enhanced productivity of infectious virion; drug screening; kits; vaccinesGoogle Patents 2006 [Google Scholar]
[41]. Deisseroth K, Airan RD, Cell line, system and method for optical control of secondary messengersGoogle Patents 2014 [Google Scholar]
[42]. Dimitrov DS, Virus entry: molecular mechanisms and biomedical applicationsNature Reviews Microbiology 2004 2(2):109-22. [Google Scholar]
[43]. Lutz A, Dyall J, Olivo PD, Pekosz A, Virus-inducible reporter genes as a tool for detecting and quantifying influenza A virus replicationJournal of Virological Methods 2005 126(1):13-20. [Google Scholar]
[44]. Li Y, Larrimer A, Curtiss T, Kim J, Jones A, Baird-Tomlinson H, Influenza virus assays based on virus-inducible reporter cell linesInfluenza and Other Respiratory Viruses 2009 3(5):241-51. [Google Scholar]
[45]. Palmer AE, Jin C, Reed JC, Tsien RY, Bcl-2-mediated alterations in endoplasmic reticulum Ca2+ analysed with an improved genetically encoded fluorescent sensorProceedings of the National Academy of Sciences of the United States of America 2004 101(50):17404-09. [Google Scholar]
[46]. Kang S, Ren D, Xiao G, Daris K, Buck L, Enyenihi AA, Cell line profiling to improve monoclonal antibody productionBiotechnology and Bioengineering 2014 111(4):748-60. [Google Scholar]
[47]. Mowafi F, Chemokines and chemokine receptors during viral infections in man: Institutionen för mikrobiologi, tumör-och cellbiologi/Department of MicrobiologyTumor and Cell Biology 2007 [Google Scholar]
[48]. Anderson NW, Buchan BW, Ledeboer NA, Light microscopy, culture, molecular, and serologic methods for detection of herpes simplex virusJournal of Clinical Microbiology 2014 52(1):2-8. [Google Scholar]
[49]. Atmar RL, Immunological Detection and Characterization. Viral Infections of HumansSpringer 2014 :47-62. [Google Scholar]
[50]. Ma JZ, Russell TA, Spelman T, Carbone FR, Tscharke DC, Lytic gene expression is frequent in HSV-1 latent infection and correlates with the engagement of a cell-intrinsic transcriptional responsePLoS pathogens 2014 10(7):e1004237 [Google Scholar]
[51]. Fan F, Day S, Lu X, Tang Y-W, Labouratory diagnosis of HSV and varicella zoster virus infectionsFuture Virology 2014 9(8):721-31. [Google Scholar]
[52]. Marks LV, The Lock and Key of Medicine: Monoclonal Antibodies and the Transformation of Healthcare 2015 Yale University Press [Google Scholar]
[53]. Portela A, Melero JA, Martínez C, Domingo E, Ortín J, A primer vector system that allows temperature dependent gene amplification and expression in mammalian cells: regulation of the influenza virus NSI gene expressionNucleic acids research 1985 13(22):7959-77. [Google Scholar]
[54]. Schägger H, Tricine–SDS-PAGENature Protocols 2006 1:16-22. [Google Scholar]
[55]. Kao J-H, Chen P-J, Lai M-Y, Chen D-S, Hepatitis B genotypes correlate with clinical outcomes in patients with chronic hepatitis BGastroenterology 2000 118(3):554-59. [Google Scholar]
[56]. Sacks G, Reproductive immunology: the relevance of labouratory research to clinical practice (and vice versa)Human Reproduction 2014 :deu325 [Google Scholar]
[57]. Viswanathan S, Keating A, Deans R, Hematti P, Prockop D, Stroncek DF, Soliciting strategies for developing cell-based reference materials to advance mesenchymal stromal cell research and clinical translationStem cells and development 2014 23(11):1157-67. [Google Scholar]
[58]. Nagy A, Jirinec T, Cerníková L, Jirincová H, Havlícková M, Large-Scale Nucleotide Sequence Alignment and Sequence Variability Assessment to Identify the Evolutionarily Highly Conserved Regions for Universal Screening PCR Assay Design: An Example of Influenza A VirusPCR Primer Design 2015 :57-72. [Google Scholar]
[59]. Barnett D, Louzao R, Gambell P, De J, Oldaker T, Hanson CA, Validation of cell-based fluorescence assays: Practice guidelines from the ICSH and ICCS–part IV–postanalytic considerationsCytometry Part B: Clinical Cytometry 2013 84(5):309-14. [Google Scholar]
[60]. Stacey GN, The Challenge of Standardization in Stem Cell Research and DevelopmentStem Cell Banking: Springer 2014 :11-8. [Google Scholar]
[61]. Franco E, Bagnato B, Marino MG, Meleleo C, Serino L, Zaratti L, Hepatitis B: Epidemiology and prevention in developing countriesWorld Journal of Hepatology 2012 4(3):74 [Google Scholar]
[62]. Yu M-W, Yeh S-H, Chen P-J, Liaw Y-F, Lin C-L, Liu C-J, Hepatitis B virus genotype and DNA level and hepatocellular carcinoma: a prospective study in menJournal of the National Cancer Institute 2005 97(4):265-72. [Google Scholar]
[63]. Kao J-H, Wu N-H, Chen P-J, Lai M-Y, Chen D-S, Hepatitis B genotypes and the response to interferon therapyJournal of Hepatology 2000 33(6):998-1002. [Google Scholar]
[64]. Koo ES, Yoo C-H, Na Y, Park SY, Lyoo HR, Jeong YS, Reliability of non-culturable virus monitoring by PCR-based detection methods in environmental waters containing various concentrations of target RNAJournal of Microbiology 2012 50(5):726-34. [Google Scholar]
[65]. Doerr H, Replacement of biologic by molecular techniques in diagnostic virology: Thirty years after the advent of PCR technology—do we still need conventional methods?Med Microbiol Immunol 2013 202(6):391-92. [Google Scholar]