Non-diphtheritic Corynebacteria are now being increasingly recognised as the causative agents of various infections. Among these organisms, Corynebacterium amycolatum is the most frequently isolated one. It has been isolated from urine, pus, catheter tips, blood, prostatic secretion, cerebrospinal fluid and sputum. However, to the best of our knowledge, there are no reports on its role in ear infections. Here, we present 12 cases of ear infection with C.amycolatum. A high index of suspicion is necessary for identification of these gram positive bacilli as they resemble other Corynebacterium species on gram stain. They have metachromatic granules which can be demonstrated by Albert’s stain and form characteristic dry, flat colonies on blood agar. These organisms are frequently resistant to ceftriaxone and imipenem. In our study, among the 12 isolates, eight isolates were resistant to ceftriaxone and four to imipenem and two were intermediately susceptible to ceftriaxone although all the 12 strains were uniformly susceptible to vancomycin. All the isolates were negative for toxA and toxB genes by PCR. Genomic sequencing of two isolates confirmed them as C.amycolatum. C.amycolatum is a relatively rare cause of pyogenic ear infections. As it demonstrates more antibiotic resistance than other similar organisms, careful identification with antibiotic susceptibility testing is required in managing these infections.
Diphtheroids, Ear infection, Resistance, ToxA and toxB genes
Case Series
There were 12 cases of Corynebacterium amycolatum which were isolated from purulent discharge from the ear in eight months (1st August 2014 to 31st March 2015). All of them presented to Otorhinolaryngology Department, Christian Medical College, Vellore, India with the symptoms of discharge from the ear [Table/Fig-1].
Demographic & clinical details of the patients
| Case | Age | Sex | Underlying co-morbidities | Ear discharge seen in | Treatment |
|---|
| 1 | 36 | Female | Skull base osteomyelitis | Right | Oral high dose ciprofloxacin for three months |
| 2 | 30 | Female | Complicated CSOM with brain abscess | Right | Drainage of brain abcess, mastoidectomy along with combination of intravenous penicillin, ceftraixone, metronidazole and gentamicin |
| 3 | 56 | Female | Diabetes mellitus | Left | Oral and topical ciprofloxacin ear drops for one week |
| 4 | 39 | Male | Chronic myeloid leukaemia | Right | Ciprofloxacin ear drops. Had relapse of discharge but resolved with combination of ciprofloxacin and betadine ear drops |
| 5 | 3 | Female | Failure to thriveRecurrent ASOM | Left | Oral amoxicillin/clavulunate and topical ciprofloxacin ear drops |
| 6 | 43 | Male | Postoperative ear impaired glucose tolerance | Left | Topical ciprofloxacin ear drops for one week |
| 7 | 49 | Female | Aplastic anaemia | Right | Topical ciprofloxacin ear drops |
| 8 | 36 | Female | CSOM with osteoma | Right | Oral and topical ciprofloxacin ear drops |
| 9 | 1 | Male | Iron deficiency anaemia | Right | Oral amoxicillin /clavulunate and topical ciprofloxacin ear drops |
| 10 | 65 | Female | Diabetes mellitus with chronic renal failure | Left | Oral and topical ciprofloxacin ear drops |
| 11 | 13 | Male | CSOM | Left | Oral and topical ciprofloxacin ear drops |
| 12 | 32 | Female | Complicated CSOM with skull base osteomyelitis | Left | High dose oral and topical ciprofloxacin ear drops |
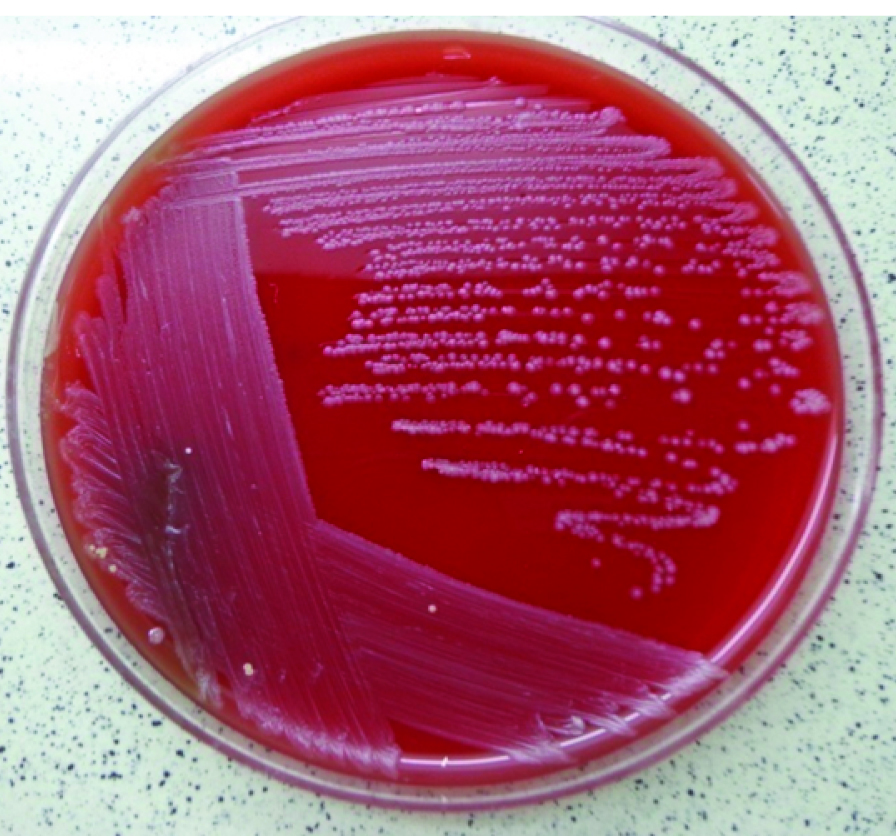
The discharge was collected using sterile swabs and sent to the Department of Clinical Microbiology for isolation and identification of the pathogens. On direct microscopy, polymorphonuclear cells with numerous Gram positive bacilli were seen which were arranged in palisades resembling the typical morphology of the diphtheroids. When special staining techniques such as Albert’s stain and Ponder’s stain were carried out on the pus, green bacilli with brown-black granules and blue bacilli with pink granules were seen, respectively. These differed from C.diphtheriae in being thicker, arrangement in palisades and having fewer granules. In contrast, C.diphtheriae are usually thin gram positive bacilli arranged in cuneiform pattern with abundant metachromatic granules. On culture, on blood agar there was heavy growth of fine colonies after overnight incubation. After 48 hours of incubation, these colonies were found to become flat, dry white colonies, which showed gram positive bacilli with granules on smears [Table/Fig-2].
Colony morphology of C.amycolatum on blood agar
The organisms were identified as Corynebacterium amycolatum by biochemical characterisation and also by the automated system Vitek 2 (Biomerieux, Marcy L Etoile, France). The organism produced catalase, used OF glucose fermentatively and fermented maltose and mannose. Methyl red production was variable. It did not reduce nitrate, decarboxylate lysine, hydrolyse esculin and lactose, sucrose, arabinose, galactose, raffinose, salicin, trehalose and xylose were not fermented.
On performing antibiotic susceptibility by minimum inhibitory concentration method according to the CLSI (Clinical Laboratory Standards Institute) 2015 guidelines it was found that all 12 isolates were uniformly susceptible to vancomycin (100%). However, resistance to ceftriaxone was seen in 8(66.7%) isolates, intermediate susceptibility in 2(16.7%) isolates. Four (33.3%) of the isolates were resistant to imipenem and 6(50%) were resistant to moxifloxacin [Table/Fig-3].
Antimicrobial susceptibility of the cases
| Antimicrobial | Vancomycin | Ceftriaxone | Imipenem | Moxifloxacin |
|---|
| Case 1 | Susceptible | Susceptible | Susceptible | Susceptible |
| Case 2 | Susceptible | Resistant | Resistant | Resistant |
| Case 3 | Susceptible | Intermediate | Susceptible | Intermediate |
| Case 4 | Susceptible | Resistant | Resistant | Resistant |
| Case 5 | Susceptible | Susceptible | Susceptible | Intermediate |
| Case 6 | Susceptible | Resistant | Resistant | Intermediate |
| Case 7 | Susceptible | Resistant | Resistant | Resistant |
| Case 8 | Susceptible | Intermediate | Susceptible | Intermediate |
| Case 9 | Susceptible | Resistant | Susceptible | Resistant |
| Case 10 | Susceptible | Resistant | Susceptible | Resistant |
| Case 11 | Susceptible | Resistant | Susceptible | Resistant |
| Case 12 | Susceptible | Resistant | Susceptible | Intermediate |
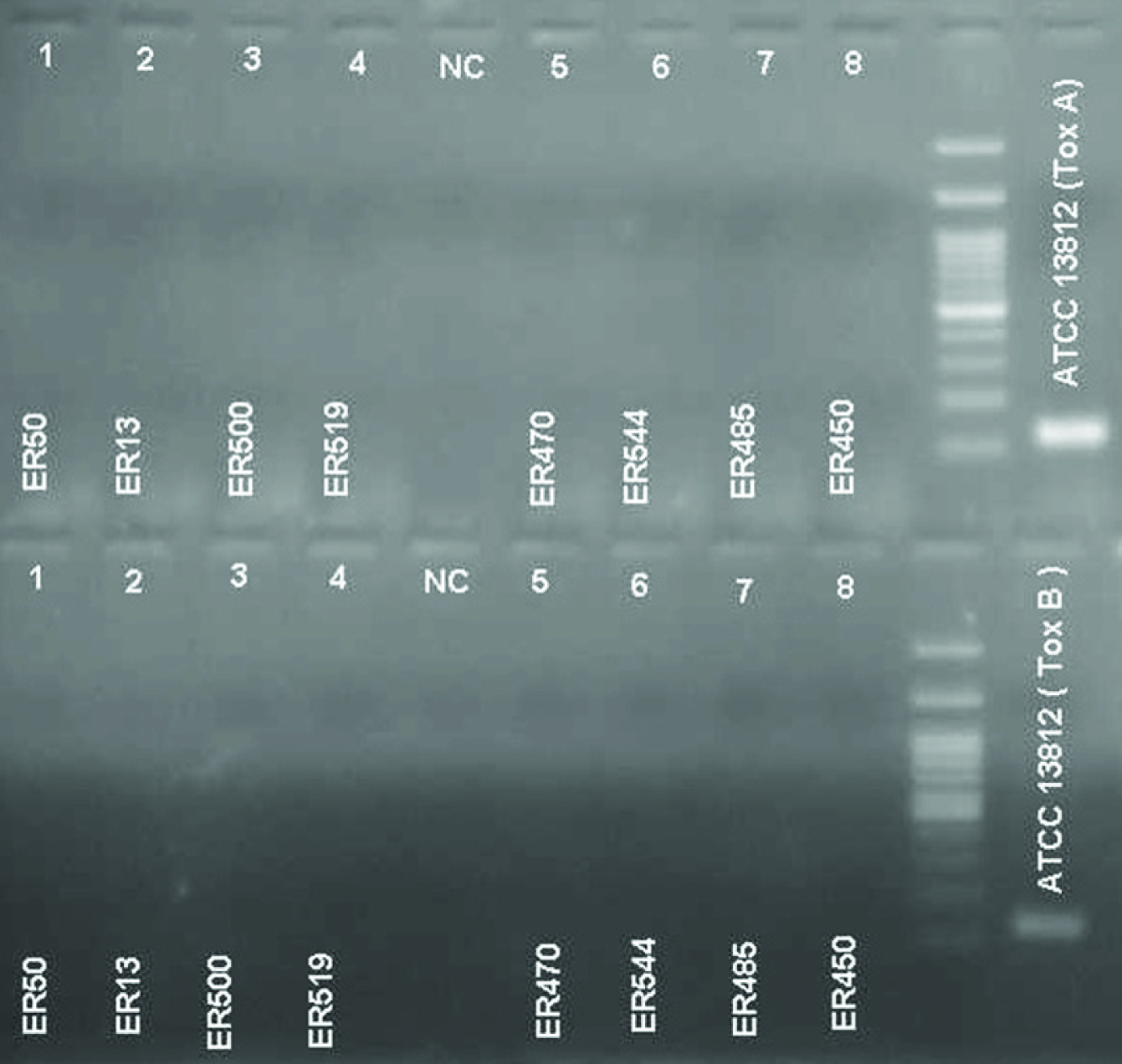
Polymerase chain reaction was negative for tox A and tox B genes of Corynebacterium diphtheriae on all the isolates, using the primer sequence as given by Mothershed et al., [1]. The primer sequences and cycling conditions are given – Subunit A (117bp) Forward – GGCGTGGTCAAAGTGACGTA, Reverse – CTTGCTCCATCAACGGTTCA and Subunit B (128bp) Forward – CGCCCTAAATCTCCTGTTTATGTT, Reverse – GTACCCAAGA ACGCCTATGGAA. Cycling conditions were 95°C for 2 min, 95°C for 20 s, 55°C for 30 s,72°C for 1 min and 72°C for 10 min for 35 cycles. It was found that none of the isolates had the toxigenicity gene implicating that it was not C.diphtheriae [Table/Fig-4].
PCR gel electrophoresis of the samples
In addition, two isolates were subjected to 16S rRNA gene sequencing for bacterial identification and they were confirmed as C.amycolatum (GenBank KR090572 and KR090573). The universal primers used for sequencing were Forward - AGAGTTTGATCCTGGCTCAG and Reverse - ACGGCTACCTTGTTACGACTT with the cycling conditions of initial denaturation 95°C for 15 min, 35 cycles of denaturation 95°C for 30 Sec, annealing 56°C for 1 min, extension 72°C for 1 min, and a final extension 72°C for 5 min (Shrimpex Biotech Services Pvt Ltd, Chennai).
Out of the 12 samples, 7(58.33%) had pure and heavy growth of C.amycolatum. Three samples had growth of C.amycolatum along with Pseudomonas aeruginosa while two had C.amycolatum with Staphylococcus aureus. However, all the samples had heavy growth of C.amycolatum.
Two samples were received for the patient case 4. Her initial sample was received in December 2014 which showed heavy and pure growth of C.amycolatum. As her ear discharge persisted even after treatment with ciprofloxacin a second sample was received in March 2015 which again showed growth of the same organism with same susceptibility pattern. After the first course of antibiotic failure, a repeat culture was sent, and the patient was treated with repeated regular ear suction clearance, thrice daily use of betadine ear drops for one month and ciprofloxacin ear drops for one more week. As it was a localised infection systemic antibiotics were not given. The ear discharge resolved after one month.
Discussion
Corynebacteria other than C.diphtheriae cause quite a few infections of the body [2]. Other Corynebacterium species like C.amycolatum, C.aurimucosum, C.glucuronolyticum, C.jeikeium, C.pseudodiphtheriticum, C.striatum, C.tuberculostearicum and C.urealyticum are isolated in the laboratory [3].
Corynebacteria are part of the normal skin flora, therefore it becomes difficult to distinguish between infection, colonization, and contamination when these organisms are isolated from purulent ear specimens. Attribution of a causal role is strengthened when polymorphonuclear leukocytes are observed in the clinical specimen along with gram positive bacilli, especially in the absence of any other pathogen [4]. In our study, all the 12 cases had the presence of polymorphonuclear cells with gram positive bacilli. Moreover, all the cases had heavy growth of the organism which is also suggestive of its role in the infection.
Corynebacterium amycolatum is a gram positive bacilli arranged in palisades. As other non-pathogenic corynebacteria resemble C.amycolatum, the microbiological diagnosis is frequently missed, if care is not taken to identify the isolate. Esteban et al., have reported that 82% of the isolates which were previously identified as Corynebacterium xerosis actually turned out to be C.amycolatum when retested [4].
C.amycolatum is associated with septicaemia, endocarditis, meningitis, septic arthritis and urinary tract infections [2]. C.amycolatum is most commonly isolated from wound swabs (61.5%), urine (14.3%), drain swabs (7.1%) [5]. However, a causative role for this organism in otitis media still remains obscure. In a study by Aalbaek et al., C.amycolatum was found to be associated with canine otitis externa [6]. We have detected 12 cases of C.amycolatum in individuals with ear infection. These patients were either immunocompromised or have had previous surgical interventions. PCR and genomic sequencing identified the organism as C.amycolatum. Out of these patients seven had pure growth of C.amycolatum while the rest had growth of other organisms. In absence of any other organism, C.amycolatum is most likely to be the causative agent of the infection. Though the outcome was not known in most of the patients, one patient showed persistence of infection as evidenced by isolation of the same agent 3 months later.
C.amycolatum is frequently resistant to many antimicrobials. C.amycolatum causing endocarditis was found to be resistant to multiple antibiotics [7]. There are reports stating that the use of broad-spectrum antibiotics led to an increase of sepsis due to the species C.jeikeium and C.amycolatum [8]. In a study done on surgical site infections it was seen that 11.2% of the infections were caused by C.amycolatum. All the corynebacteria were multidrug resistant, and 51.8% of isolates were sensitive to only gatifloxacin and vancomycin [9]. In another study by Reddy et al., it was demonstrated that these organisms were resistant to penicillin, erythromycin, and clindamycin but susceptible to vancomycin, linezolid, and tigecycline [2]. In the 12 cases, we found resistance to ceftriaxone in 66.7% and to imipenem in 33.3% cases. Hence, it is important that an antimicrobial susceptibility testing be performed on each isolate to determine the treatment modality in case of infection with these organisms.
Conclusion
This report provides preliminary information that C.amycolatum could be an ear pathogen especially in patients who are immunocompromised or have had surgical interventions. It is likely that many infections are missed as this organism is often mistaken as normal skin flora or a contaminant. Our findings suggest that C.amycolatum could be a potential pathogen especially when it is isolated as a pure and heavy growth with surrounding inflammatory reaction. In addition, this organism is more resistant to many of the commonly used antimicrobials as compared to other Corynebacteria. Further prospective studies are needed to confirm the role of C. amycolatum as a pathogen in the ear.
[1]. Mothershed EA, Cassiday PK, Pierson K, Mayer LW, Popovic T, Development of a Real-Time Fluorescence PCR Assay for Rapid Detection of the Diphtheria Toxin GeneJ Clin Microbiol 2002 40(12):4713-19. [Google Scholar]
[2]. Reddy BS, Chaudhury A, Kalawat U, Jayaprada R, Reddy G, Ramana BV, Isolation, speciation, and antibiogram of clinically relevant non-diphtherial Corynebacteria (Diphtheroids)Indian J Med Microbiol 2012 30(1):52-57. [Google Scholar]
[3]. Cazanave C, Greenwood-Quaintance KE, Hanssen AD, Patel R, Corynebacterium prosthetic joint infectionJ Clin Microbiol 2012 50(5):1518-23. [Google Scholar]
[4]. Esteban J, Nieto E, Calvo R, Fernández-Robals R, Valero-Guillén PL, Soriano F, Microbiological characterization and clinical significance of Corynebacteriumamycolatum strainsEur J ClinMicrobiol Infect Dis Off Publ Eur Soc Clin Microbiol 1999 18(7):518-21. [Google Scholar]
[5]. Zalas P, Mikucka A, Gospodarek E, Occurrence of Corynebacteriumamycolatum strains in clinical specimensMed Dośw Mikrobiol 2004 56(2):139-45. [Google Scholar]
[6]. Aalbæk B, Bemis DA, Schjærff M, Kania SA, Frank LA, Guardabassi L, Coryneform bacteria associated with canine otitis externaVet Microbiol 2010 145(3-4):292-98. [Google Scholar]
[7]. Knox KL, Holmes AH, Nosocomial endocarditis caused by Corynebacteriumamycolatum and other nondiphtheriaecorynebacteriaEmerg Infect Dis 2002 8(1):97-99. [Google Scholar]
[8]. Riegel P, Bacteriological and clinical aspects of corynebacteriumAnn BiolClin (Paris) 1998 56(3):285-96. [Google Scholar]
[9]. Rizvi M, Rizvi MW, Shaheen null, Sultan A, Khan F, Shukla I, Emergence of coryneform bacteria as pathogens in nosocomial surgical site infections in a tertiary care hospital of North IndiaJ Infect Public Health 2013 6(4):283-88. [Google Scholar]