The objectives of the present study were to apply the molecular methods for the species identification of the Acinetobacter spp. clinical isolates and to determine their antibiotic susceptibility patterns.
Materials and Methods
Bacterial isolates
A total of 197 non-duplicate isolates from a wide range of clinical samples were collected from Golestan and Imam Khomeini teaching Hospitals in Ahvaz and Sina Hospital in Tehran from November 2011 to January 2013. Clinical sources of the isolates were blood, endotracheal, urine, wound, sputum, exudates, percutaneous endoscopic gastrostomy (PEG), pleural fluid, cerebrospinal fluid (CSF), and soft tissue, though the majority of strains were isolated from endotracheal tube and blood. The isolates were kept in Trypticase Soy Broth (Merck, Germany) containing 20% glycerol at -80°C until use. Conventional phenotypic tests including growth on MacConkey agar (Merck, Germany), sugar fermentation, motility, catalase, oxidase, and other standard recommended tests were used to identify the genus of the isolates [15].
Antimicrobial susceptibility testing
Antimicrobial susceptibility testing was performed by agar disc diffusion method according to Clinical and Laboratory Standards Institute (CLSI) guidelines [16]. The following antimicrobial agents were tested: ciprofloxacin (30μg), piperacillin (100μg), gentamycin (10μg), amikacin (30μg), trimethoprim sulfamethoxazole (25μg), meropenem (10μg), piperacillin tazobactam (100/10μg), ceftazidime (30μg), ceftriaxone (30μg), polymyxinB (20μg) and tetracycline (30μg) {MAST Co., UK}. A. baumannii ATCC 16906 was used as control strain. Determination of MDR and extensively drug resistance (XDR) was based on the CLSI criteria reported elsewhere [17].
Molecular identification
DNA extraction
All the isolates were grown on Muller Hinton Agar (Merck, Germany) and were incubated for 48 hours at 37°C under aerobic conditions and the bacterial colonies were then used for DNA extraction. The boiling method described by Higgins et al., [18], was used to extract the DNA from the bacterial colonies.
Fingerprinting of the 16S rRNA gene by PCR-RFLP analysis for strain clustering
Amplification of the 16S rRNA gene (1500 bp) was performed by using universal primers 27F (5’-AGAGTTTGATYMTGGCTCAG-3’) and 1525R (5’-AGAAAGGAGGTGATCCAGCC-3’) and PCR protocol as described elsewhere [14,19]. The PCR products were electrophoresed on a 2% agarose gel containing 0.5 μg/ml ethidium bromide (Qiagen, USA) to determine the size of the product. Both negative and reagent controls were included in each PCR run. The product was photographed by using the gel documentation system (UV Tech, UK). All PCR products were then digested by Hae III and Tag I restriction enzymes (Fermentas, Canada). Enzymatic digestion of the PCR products was performed according to the manufacturer's instructions. In brief, 10 μl of the amplicon was treated separately with 2 units of TaqI or Hae III enzymes plus 10 μl corresponding buffer and were overnight incubated at 37oC. The digested reactions were then placed at 65°C for 30 minutes to avoid further enzymatic digestion. Ten microliters of digestion products were separated by electrophoresis on a 3% agarose gel containing 0.5 μg/ml ethidium bromides. A. baumannii ATCC16906 was used as a positive control. Restriction patterns were analysed visually comparing with a 1000 bp DNA size marker, to group the patterns according to a similarity coefficient using the UPGMA method, allowing a 5% level of discrimination between the bands.
Nucleotide sequence determination of rpoB and 16S rDNA genes and phylogenetic analysis
One isolate from each cluster was selected randomly for further identification by sequencing of 16S rDNA and rpoB. The PCR products were sequenced using 27F, 1525R and 907R (5’-CCGTCAATTCMTTTRAGTTT-3’) with an ABI 3100 genetic analyser. To determine the sequencing of PCR products, commercial kit of Dye Terminator Cycle Sequencing Genmeo Lab TM (Biometra Co., Germany), according to the manufacturer’s instructions was used.
Zone 1 of the rpoB gene (350bp) was amplified from randomly selected isolate of each group using the primers Ac696F (5’-TAYCGYAAAGAYTTGAAAGAAG-3’) and Ac1093R (5’-CMACACCYTTGTTMCCRTGA-3’) as described elsewhere [14]. The PCR products were sequenced using Ac696F and Ac1093R.
Multiple alignments, sequence similarities of the 16S rDNA and rpoB gene were determined with all existing sequences of type strain of Acinetobacter retrieved from GenBankTM database using the jPhydit program [20].
The phylogenetic relationship between sequences of the 16S rDNA and rpoB genes was estimated by the neighbor-joining method. After the construction of phylogenetic trees, bootstrap replicates were performed to estimate their node reliabilities using the MEGA 4 program.
The GenBank accession numbers for the sequences of the 16S rRNA and rpoB genes of the representative isolates are KM281495-KM281506 and KM668180-KM668191 respectively.
Results
The clinical isolates of Acinetobacter belonged to 130 male (66%) and 67 female (34%) patients with an age range of 2 to 84 years. The majority of the isolates were from endotracheal tubes (95 isolates, 48.2%), followed by ulcer discharge (26 isolates, 13.2%) and urine (27 isolates,13.7%). The results from susceptibility testing showed that 159 (81%) Acinetobacter isolates were MDR and 26 (13%) were XDR. The rate of antibiotic susceptibility was only 6% (12 isolates). The highest antibiotic resistance among Acinetobacter isolates were equally (96%) seen against ceftriaxone, piperacillin and piperacillin tazobactam, while polymyxin B was the most sensitive antibiotic (49.2%) in present study [Table/Fig-1].
Antibiotic susceptibility patterns of clinical isolates of Acinetobacter spp.
| Number (Acinetobacter) | Isolates | AMK | CIP | CTZ | PMB | MRP | TMP-SMX | CTX | TZP | PIP | GEN | TCN | MDR Status |
|---|
| 57,136,113, 82 | A.pittii | S | S | I | S | S | S | I | S | I | R | S | _ |
| 25,34,85,148 | A.schindleri | R | R | R | S | I | R | R | R | R | R | R | XDR |
| AC99,2, 17,28 | A.calcoaceticus | S | S | S | S | R | S | R | R | R | S | S | MDR |
| 70,104,117,129 | A.gen.sp. 16 | R | R | R | S | R | R | R | R | R | R | I | MDR |
| 79,96,108,120 | A.gen.sp. 9 | R | R | R | I | R | R | R | R | R | R | I | MDR |
| 35,12,20,22,23,52,48,49 | A.baumanii | S | S | S | S | S | S | S | S | S | S | S | _ |
| 60,65,95*161,66, 67,69,80,81 | A.baumanii | R | R | R | S | R | R | R | R | R | R | S | XDR |
| 51,55,58,36,37,54,61,62,46,162 | A.baumanii | R | R | R | S | R | I | R | R | R | S | I | MDR |
| 41, 45 ,50,9 ,14,15, 32,123,71 | A.baumanii | R | R | R | R | R | R | R | R | R | R | I | MDR |
| 26,29,40,53,64,76,89,101,125,152,4,18,6,59,24,72,73,74,84,86,98,19,30,31,42 ,44 | A.baumanii | R | R | R | S | I | R | R | R | R | R | I | MDR |
| 1,8,11,16,27,38,75,88,100,112, 124,135 151,91,93,94,103,105,115,118,127,130,138,139,140,156,7,158,168,107,109,110,166,119,121,122,131,132,15,133,134,143,144,149,159,160,162,170,171,163,77,90,102,114,153,165,116,167,126,137 | A.baumanii | R | R | R | R | R | I | R | R | R | R | R | MDR |
| 313,470,79,415,553 527,27,475,545 | A.baumanii | R | R | R | S | R | R | R | R | R | S | R | XDR |
| 99,384,169,339,379 | A.lwoffii | R | R | I | I | R | R | R | R | R | I | R | MDR |
| 36,505,25,262 | A.ursingii | R | S | S | S | R | R | R | R | R | R | R | MDR |
| 20,476,420,546,550,670 | A.johnsonii | R | R | R | S | R | I | R | R | R | R | I | MDR |
| 227,106,233,406 | A.parvus | S | R | R | S | I | R | R | R | R | I | I | MDR |
| 218,49,56,121 | A.gen.sp. 3 | R | R | R | R | R | R | R | R | R | R | R | XDR |
| 544,59,502,409 | A.gen.sp.16 | S | R | R | S | I | R | R | R | R | R | I | MDR |
| 544,59,502,409 | A.genompsp.16 | S | R | R | S | I | R | R | R | R | R | I | MDR |
| 555,67,127,552 | A.girmontii | S | R | R | I | R | I | R | R | R | S | I | MDR |
| 364,255,259,319,336 | A.calcoaceticus | S | I | S | S | R | I | R | R | R | I | I | MDR |
| 383,26,41,76,469,509 | A.gen.sp. 13 TU | S | R | R | R | I | R | R | R | R | S | I | MDR |
Isolates were designated susceptible (S), intermediate (I), or resistant (R) according to antibiotic breakpoint guidelines of the CLSI for Acinetobacter spp. Antibiotic abbreviations: AMK: Amikacin, CIP: Ciprofloxacin, CTZ: Ceftazidime , PMB: Polymyxin B , MRP: Meropenem , TMP-SMX: Trimethoprim Sulfamethoxaxole, CTX: Ceftriaxone , TZP: Piperacillin tazobactam , PIP: Piperacillin, GEN: gentamicin, TCN: Tetracycline.
The isolates showed several RFLP patterns which were grouped into 13 clusters designated from A to M. The majority of isolates were classified in A (131 isolates, 66.5%) [Table/Fig-2,3]. Most of clinical isolates can be confidently identified with high bootstrap values using 16S rRNA gene sequencing. However based on 16S rRNA gene, the identity of the isolate 555T was uncertain due to high similarity of A. grimontii and A. junii. Similarly on the basis of 16S rRNA gene, the isolates 218T and 364T were unidentifiable due to identical 16S rRNA sequence of genomic species (Gen.sp.) 3 and A. calcoaceticus.
Restriction patterns and clustering of clinical isolates of Acinetobacter spp. by 16S rRNA RFLP.
| Number of isolates | Hae III | Tag I | RFLP type |
|---|
| 131 | 100/170/220/250/300/400 | 100/220/300/320/480 | A |
| 4 | 100/220/250/350/400 | 220/350/370/470 | B |
| 4 | 50/120/250/300/400 | 200/310/400/600 | C |
| 8 | 100/170/220/250/300/400 | 120/250/320/480/600 | D |
| 4 | 100/170/220/250/300/400 | 70/180/420/610 | E |
| 9 | 100/170/220/250/300/400 | 70/180/320/610 | F |
| 10 | 70/180/230/400 | 90/160/400/520 | G |
| 5 | 80/170/200/250/300/390 | 220/300/320/450 | H |
| 4 | 90/170/220/250/470 | 100/290/300/320/550 | I |
| 6 | 100/220/250/300/550 | 50/280/300/320/450 | J |
| 4 | 170/210/250/400/630 | 90/230/300/320/400 | K |
| 4 | 170/320/450/510 | 80/120/300/320/390 | L |
| 4 | 170/210/250/420/680 | 90/210/300/320/400 | M |
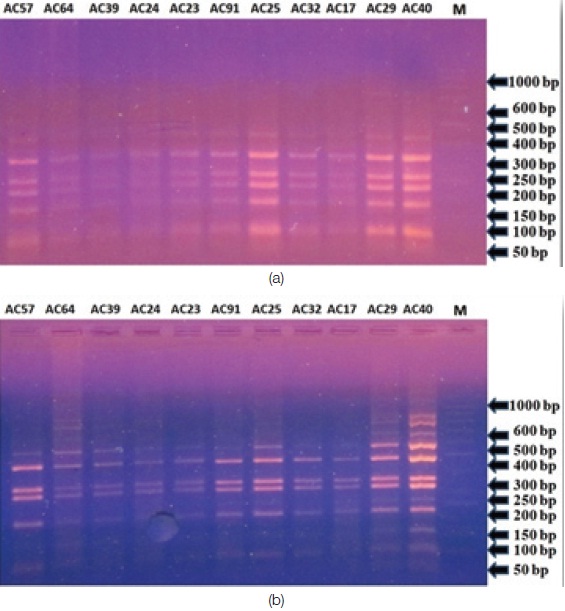
Enzymatic digestion of 16S rRNA gene PCR products with (a) Hae III; (b) Tag I restriction enzymes with subsequent separation of the digested products by 3% agarose gel electrophoresis.Lanes: M,1000-bp DNA size marker; 1-11 Acinetobacter isolates.
Sequence analysis of zone 1 of the rpoB gene for representative isolates from each cluster in RFLP grouping, identified most of the isolates to species level with high similarity. Based on rpoB gene, clinical isolate 555T showed 100% and 99% similarity to those of A. grimontii. and Gen. sp. 5 (A. junii) respectively. Based on rpoB gene, the isolates 218T and 364T were identifiable to species level confidently of Gen. sp.3 A. calcoaceticus.
Identification by concatenation of 16S rRNA and rpoB confirmed the identity of clinical isolates of Acenitobacer to species level. Accordingly, the isolate 555T assigned as A. grimontii due to 100% similarity to A. grimontii [Table/Fig-2]. Moreover, this isolate showed 98.64% (24 nucleotide mismathes) to A. junii. On the other hand, the identity of the isolates 218T and 364T was confirmed as Gen. sp. 3 and A. calcoaceticus respectively [Table/Fig-4].
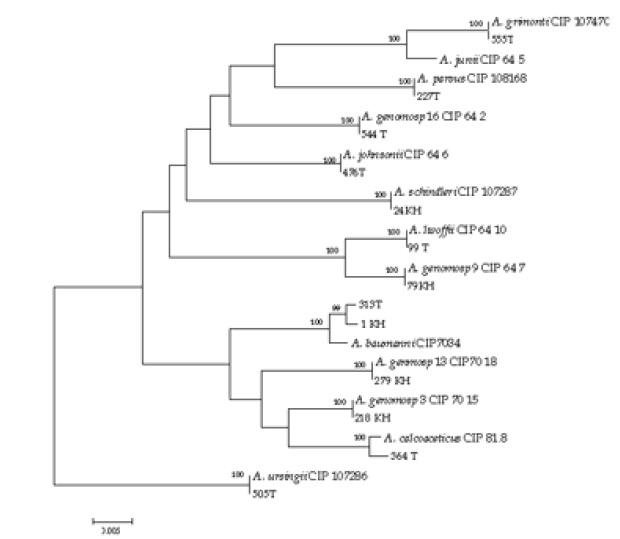
Phylogenetic tree of the genus Acenitobacter computed from the concatenation of 16S rRNA and rpoB gene sequences by the Neighbor joining method and Kimura 2-parameter (K2P) model as the substitution model. Percentages at nodes represent bootstrap levels supported by 1000 re-sampled datasets. Bar 0.02 substitutions per nucleotide position.
Abbreviations: T: Tehran isolate; KH: Khuzestan isolate
Discussion
Acinetobacter spp. and mainly A. baumannii are associated with hospital outbreaks worldwide. These organisms are particularly problematic due to the large number of MDR strains that have become endemic in hospital settings which is a growing concern in many countries [21,22].
In the present study, A. baumannii was the most prevalent (66.5%) Acinetobacter species. Several studies have investigated the distribution of Acinetobacters in clinical specimens at the species level, and considerable differences in outcome have been reported. In a recent report by Lee et al., [23], about 80% of their Acinetobacter isolates were identified as A. baumannii, which was higher than our findings. This rate was much lower in the study of Karah et al., [12], as 24% and in the study of Boo et al., [24], as 22%. In the latter study, A. pittii was the most frequent species found, representing 39% of their 114 isolates, whereas only 4 isolates in our study were identified as A. pittii, which the difference is considerable. This inconsistency may be explained either by the use of different interval surveillance cultures in their study or by the use of different methods for species identification. However, whether these differences are due to geographic, methodological, or epidemiological differences between the studies is hard to say.
Our finding represented extremely high rate of MDR as 81% and lower rate of XDR as 13%. This was in line with a report by Joshi et al., with more than 75% MDR among their Acinetobacter isolates [25]. Similarly a recent report from Pakistan represented higher multidrug resistance as 96.6% among their tested Acinetobacter strains [9]. Although there are some discordant reports with much lower rate of MDR as 5.6% and 7.7% respectively [26,27]. A likely explanation for the relatively low level of resistant Acinetobacter isolates in latter studies is probably the restricted use of antibiotics, which is in line with the practice regarding antimicrobial therapy in their countries, the strict isolation of patients with MDR isolates, and the immediate beginning of infection control measures when several patients are infected with the same strain.
Identification to species level within the genus Acinetobacter is often problematic. Currently Acinetobacter species are defined on the basis of a battery of phenotypic tests and several genotypic methods [12]. Among the latter, 16S rRNA gene sequencing is one of the most commonly used for bacterial identification [22]. However, the main limitation of 16S rRNA gene sequencing is due to being too conserved and is not polymorphic enough to differentiate all Acinetobacter spp. [28]. Other protein-encoding genes, including recA, rpoB, gyrB, have recently been used for the identification of Acinetobacter isolates. Among them, rpoB gene due to variability in its size in different bacterial species, comprises priority for typing subspecies [12,29,30]. In the current study, the isolates were subjected to phenotypic and sequence based identification to access the species spectrum of clinically important species of Acinetobacter.
The obtained results revealed that although the 16S rRNA is able to identify most isolates to species level, however as previously stated, because of low polymorphisms, it is not suitable genetic marker for identification. The isolates 555T, 218T and 364T were poorly discriminated by the 16S rRNA gene sequence. Identity of isolate 555T was unclear by rpoB sequencing as well. Although, simultaneous application of both 16S rRNA and rpoB techniques, was able to identify the randomly selected isolates to species level. Our finding showed 98.64% similarity between A. grimontii and A. junii. These are the species which previously were shown to be identical based on the complete rpoB gene sequences [18]. Other pair species showed less similarities by combination techniques and could not assigned as the identical species in pair wise comparisons.
According to the reports by other investigators, the discriminating power of 16S rRNA gene sequencing is known to be lower than that of rpoB gene sequencing [14,31]. Although the rpoB significantly improved the resolution of species identification and complemented the phylogeny of genus Acinetobacter based on 16S rRNA gene sequences, in our study, using the concatenated sequence analysis of 16S rRNA and rpoB of representative isolate of each RFLP pattern, facilitated molecular identification for clinical isolates [Table/Fig-4]. These data clearly showed that identification based on single technique is not suitable for reliable identification and combination of 16S rRNA and rpoB genes analysis markedly improved the ability of the sequence based identification.
Conclusion
Sequence analysis of the 16S rRNA and rpoB spacer simultaneously was able to do identification of Acinetobacter spp. to species level. A. baumannii was identified as the most prevalent species with high antibiotic resistance. Other species showed lower frequencies ranged from 4 to 9 strains.
Isolates were designated susceptible (S), intermediate (I), or resistant (R) according to antibiotic breakpoint guidelines of the CLSI for Acinetobacter spp. Antibiotic abbreviations: AMK: Amikacin, CIP: Ciprofloxacin, CTZ: Ceftazidime , PMB: Polymyxin B , MRP: Meropenem , TMP-SMX: Trimethoprim Sulfamethoxaxole, CTX: Ceftriaxone , TZP: Piperacillin tazobactam , PIP: Piperacillin, GEN: gentamicin, TCN: Tetracycline.