The infection can be transmitted by consumption of contagious oocysts orally from water, soil or vegetables, by transplacental transmission of tachyzoites or by tissue growths contained in crude or half-cooked meat. By tachyzoites, infection may transmitted by organ transplantation, blood transfusion or by unpasteurised goat milk [4,5].
The final diagnosis of toxoplasmosis is fulfilled by the isolation of the parasite from patients [10]. Serological tests like Sabin-Feldman dye test, the Indirect Haemagglutination Assay (IHA) and Enzyme-Linked Immunosorbent Assay (ELISA) are generally used for diagnosis of T. gondii infection in animals and humans [10]. Molecular methods depend on PCR for the specific detection or analysis of T. gondii DNA [11]. Molecular techniques have become crucial and reliable tools [12], in diagnosis as well as in the understanding of the epidemiology of T. gondii [12].
LAMP is a new diagnostic technique that amplifies the target nucleic acids and has high sensitivity, specificity, efficiency, and rapidity under isothermal conditions. It has been already evaluated for the detection of Cryptosporidium parvum [13], Toxoplasma oocysts in water [14], Giardia lamblia [15], in other areas in Parasitology [16]. Notomi T. et al., firstly developed LAMP [17], and the method uses a DNA polymerase called Bst polymerase with displacement activity and a set of four specially designed primers that recognise a total of six distinct sequences of the target DNA. Urine samples are useful and non-invasive procedure for the diagnosis of T. gondii antigens during acute infection [18].
Materials and Methods
The present cross-sectional study was done in the Northern border area, Saudi Arabia between 2016 and 2018 in faculty of pharmacy, Northern border university, the samples were toxoplasmosis positive cases [19] confirmed by ELISA Kits (UDI EM127 Toxoplasma IgM, Dammam, KSA) and (UDI D91040 Toxoplasma IgG, Dammam, KSA) from the United Diagnostic Industries company. The test protocol was done as indicated by the manufacturer’s directions. Ultimately, 11 samples each of blood and urine from the same participant were used in the current study for the LAMP detection.
The study was carried out as directed by the Helsinki declaration, and approved by the local bioethics committee, in the Northern Border University Arar, kingdom of Saudi Arabia, (Approval No.4/37/A). Approval number (1/38) was also obtained from center of medical education and research, general directorate of health affairs in the northern border region, Ministry of Health, KSA. Approval was also given from the administration of Rafha Central Hospital. This study was a part of previously published study [19], which has been granted an ethical approval before. Written and verbal consent were taken from the participants.
DNA Extraction from Blood Samples
A solution based genomic DNA purification from a whole blood, commercial kit (Jena Bioscience GmbH, Germany. LOT # 112.811) was used according to the manufacturer’s instructions.
DNA Extraction from Urine Samples
FitAmp™ Urine DNA Isolation Kit (© Epigentek Group Inc. USA. LOT # 605172) was used according to the manufacturer’s instructions. This kit simply applies the desired DNA isolation buffer to urine sediments. After treatment with DNA digestion buffer, the DNA was easily recovered in 8-20 μL by specially designed Fast-Spin Columns.
Loop-Mediated Isothermal Amplification (LAMP) Assay
This part was done in the laboratories of the Department of Parasitology, Faculty of Medicine, University of Malaya, Kuala Lumpur, Malaysia.
The FIP, BIP, F3, and B3 primers involved in this study were adapted from Lau YL et al., [20], and described in [Table/Fig-1]. Two genes included in this study were B1 and SAG2 genes of T. gondii (GenBank accession number AF179871 and M33572).
The nucleotide sequence of Lamp Primers used for this study.
| Gene | Primer | Primer Sequence | bp | Target |
|---|
| B1-LAMP | B3 | 5’ACGTGACAGTGAAGAGAGGA-3’ | 215 | T. gondiiB1 gene (GenBank accession no. AF179871) |
| F3 | 5’CAGATGTGCTAAAGGCGTCA-3’ |
| BIP | 5’TGTTCGCTGTCTGTCTAGGGCAGGTGGTCGACTTCATGGGA-3’ |
| FIP | 5’AGGCGGAACCAACGGAAATCCTTGCTGTTCTGTCCTATCGC-3’ |
| SAG2-LAMP | B3 | 5’ GTAGCAGGACCTTTCGCG -3’ | 196 | T. gondiiSAG2 gene (GenBank accession no. M33572) |
| F3 | 5’ GCGCAACGAAGACTGTTGA-3’ |
| BIP | 5’ TGGCAAGGAATGCACAGACTCGGACCTTAGCTGTCAAGACCG -3’ |
| FIP | 5’ CGCCACTGGGACTGATGGTTAGTGCACCCTCCAGTGGTTC -3’ |
FIP: Forward inner primer; F3 Primer: Forward outer primer; BIP: Backward inner primer; B3 Primer: Backward outer primer
A 25 μL Reaction Mixture (RM) consisted of 1.6 μM FIP and BIP primer, 0.2 μM F3 and B3 primer, 12.5 μL of 2X RM (provided in the kit), 1 μL of Bst Enzyme, 0.35 μL of sterile dense water, 0.15 μL of Hydroxynaphthol Blue Dye (HNB) and 2 μL of DNA extracted from both urine and blood samples were used as template. The RM was carried out in a Loopamp real-time turbidimeter (LA-320, Teramecs, Co., Ltd., Japan) at 65°C for 60 minutes and inactivated for 2 minutes at 80°C. Five microliters of the end product were analysed by electrophoresis with 1.5% agarose gel and photographed by gel documentation system UV.
Analytical Sensitivity and Specificity of the LAMP Method
To set the analytical sensitivity of the LAMP assay, ABI Toxoplasma gondii (RH strain) quantitated DNA (positive control), was diluted from 16000 copies to 2 copies per microliter in distilled water and mixed with three genomic DNA of clinical bacterial isolates of (Klebsiella pneumonia, Escherichia coli, and Pseudomonas aeruginosa), then LAMP reaction for the mixed samples and positive control was performed.
Bacterial Isolates
Three bacterial isolates were obtained from the microbiology section, Rafha Central Hospital after identification using API 20E system (Biomerieux). A confirmatory identification of microorganisms was carried using the Vitek 2 compact system instrument, in the Department of Basic Health Sciences, Faculty of Pharmacy, Northern Border University, Saudi Arabia.
DNA Extraction from Bacterial Samples
A solution based, genomic DNA purification from bacterial culture, commercial kit (Jena Bioscience GmbH, Germany, LOT# 114. 158) was used according to the manufacturer’s instructions.
DNA Sequencing of LAMP Products
For confirmation of the specificity of the LAMP Products, conventional PCR method was applied using outer primers B3 and F3 as described by Lau YL et al., [20].
The primer F3 was used for PCR amplification, the reaction was performed in 25 μL of RM containing 5 μL of PCR buffer (10X), 5 μL (200-μmol concentration) of each dNTP, 4 μL (200 nmol concentration) (each) of primers B3 and F3, and 0.25 μL (2.5 U) of AmpliTaq Gold polymerase (Applied Biosystems, Darmstadt, Germany). Initial denaturation was done at 95°C for 10 minutes, followed by 35 cycles of denaturation (45 seconds at 95°C), annealing (60 seconds at 60°C), and extension (90 seconds at 72°C). Five-microliter aliquots of the PCR products were subjected to electrophoresis on a 1.5% agarose gel and visualised under UV light.
Purification of the PCR products was performed with a QIAquick PCR purification kit (Qiagen), using the Microcentrifuge protocol, according to the manufacturer’s instructions. PCR products were sent for sequencing with SAG2 F3 primer to (NHK Bioscience Solution, Malaysia).
Statistical Analysis
Data were analysed using SPSS (SPSS Inc. Chicago, IL. USA) version 20.0. Descriptive data were presented as frequencies and percentages.
Results
The mean age (years) ± standard deviation of the study participants was 29±7. The age of the study participants ranges between 16 and 45 years. Most women participated in this study were in the age group (26-30 years), the less group were (>40 years).
LAMP Reaction
Twenty two samples were checked by LAMP. The samples were (11) blood IgG positive and their corresponding urine samples taken from the same participant.
The data in [Table/Fig-2] indicated that, LAMP results were negative in blood samples 3, 4, 6. However, for the same participants, urine samples results were positive. Sample number one was positive in blood and negative in urine.
Electrophoresis of SAG2-LAMP Products. a) Lanes: 1 to 9, DNA extracted from blood samples. b) Lanes: 11 to 19, DNA extracted from urine samples. Lanes 1, 2, 5, 7, 8, 9, 10, 12, 13, 14, 15, 16, 17, 18 and 19 were positive. Lanes 3, 4, 6, and 11 were negative. Lanes: 10 and 20, ABI Toxoplasma gondii (RH strain) mixed with DNA extracted from (Klebsiella pneumonia, Escherichia coli, and Pseudomonas aeruginosa). Lanes: M, DNA Ladder (1 Kb).
one sample was not loaded in the electrophoresis apparatus
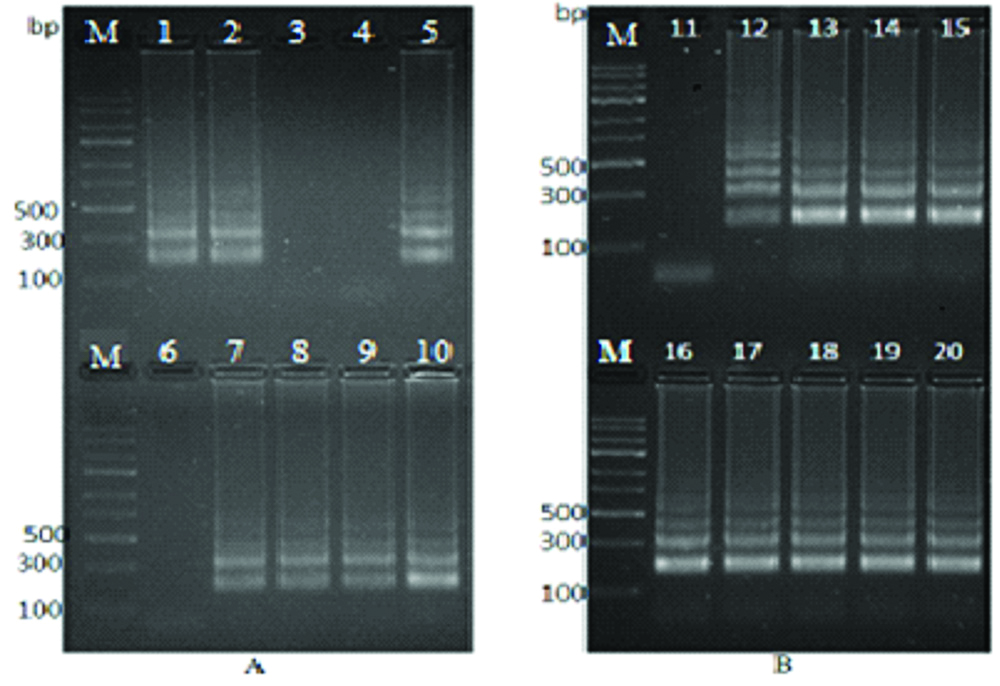
It was apparent from [Table/Fig-2] that, positive LAMP reactions produced a typical run of multiple bands on the agarose gel, indicating that stem-loop DNA with inverted repeats of the target sequences was created.
Analytical Sensitivity and Specificity of the LAMP Method
In [Table/Fig-2], lane No. 10 and 20, only the T. gondii DNA was amplified which was the target DNA, whereas no amplification was observed in the mixed bacterial DNA.
DNA Sequencing of LAMP Products
After PCR amplification of the LAMP product and sequencing, five urine samples showed 99% similarity to T. gondii isolate 66_altSAG2_W_III SAG2 gene.
Discussion
Several PCR techniques have been continuously developed for the diagnosis of toxoplasmosis using diverse clinical specimens. In this study, LAMP was used successfully to amplify Toxoplasma DNA based on the SAG2 and B1 gene, these genes were selected for their recurrent use to detect toxoplasmosis from clinical samples [20,21].
A number of reports proved the higher sensitivity of LAMP for the detection of T. gondii than conventional PCR targeting B1 and/or RE 529 bp fragment of T. gondii genome [21-24]. Another specific gene targets of T. gondii DNA was also performed using a LAMP, these were SAG1-LAMP, SAG2-LAMP, B1 gene, GRA1 gene, and TgOWP [14,20,25,26]. Up to now LAMP was used successfully for the detection of T. gondii in different animals, including sheep [23], mice [25], and pigs [27]. In humans, two published articles utilised effectively the LAMP technique [20,28].
The amplification of T. gondii was successful at 65°C for 60 minutes using the LAMP protocol in this study, there is a similarities in the temperature degree and the time required for the technique between the present study and those described by Lau YL et al., Fallahi S et al., and Sun XM et al., [20,24,28]. The high sensitivity of LAMP is because of the four primers used in the reaction. The primers target six distinct internal regions in the target DNA, and the autocycling amplification can produce very large DNA fragments of distinct sizes [29].
In LAMP procedure, DNA could be amplified using a simple water bath or heat blocker due to its isothermal conditions. Consequently, the LAMP assays could potentially be used for tests in the field. This characteristic feature not just result in higher output, but it also prevent the use of PCR machine and long cycles of different temperatures, and ultimately saving the time consumed by thermal changes for one experiment. In this study Loopamp real-time turbidimeter was used for the LAMP reaction, this is another advantage because the amplification was easily detected by visualising the turbidity of the RM even with or without the addition of dye [17,30]. This visual inspection diminishes the need for gel electrophoresis, which effectively reduces the time to analyse such results.
Nevertheless, there is a high risk of contamination, especially when opening the reaction tube in the laboratory or for the addition of the dye. Several methods were developed for obtaining better results and avoid opening the tubes, these comprise measurement of the turbidity [30], or colour change by using hydroxy naphthol blue or other chosen metal indicators [31], entrapping the fluorescent compound into the wax beads that melt at 80°C and released it in the last step of the reaction [32]. To minimise and reduce the chances of contamination, gloves should be changed frequently, implementation of sterile pipetting techniques during the assay process [20,24].
The specificity of the LAMP assay was confirmed in this study by assessment of the bacterial DNA samples and T. gondii DNA; positive amplification results were obtained only for T. gondii DNA. Moreover, in this study the specificity of the LAMP primers used for the T.gondii was proved by a conventional PCR with the outer primer F3.
Sequencing and alignment of the purified PCR products showed that the obtained partial sequences were identical to the corresponding B1 and SAG2 reported in GeneBank, this confirmed that the LAMP primers of the two genomic targets are highly specific for the detection of T.gondii.
This study showed higher sensitivity and specificity of the LAMP assay for the detection of T. gondii, these findings are similar and confirmed by previous studies which concluded the LAMP method based on the RE, SAG1, and B1 gene had a greater sensitivity than the conventional PCR based on the same genomic targets for the detection of T. gondii DNA [20-22,24,25].
The molecular detection of pathogens from urine samples for non-genitourinary infections has been successfully reported in tuberculosis [33], Leishmaniasis [34], and Malaria [35], and Toxoplasmosis [18,25].
In this research, the LAMP method successfully detects T. gondii DNA in (63.6%) of the blood samples, while 10 samples (91%), were positive using DNA extracted from urine samples. These results suggest the higher sensitivity of urine for the diagnosis, however, no data was found to use human urine for molecular diagnosis using the LAMP method, since it has been proved as accurate diagnostic tool for chronic toxoplasmosis infection in animals [24]. On the other hand, techniques other than LAMP such as traditional and membrane-based ELISA systems were used for the detection of Toxoplasma antigens and anti-Toxoplasma antibodies in urine taken from human patients [36,37].
The findings of this research suggests that, detection of the T. gondii DNA in urine may be attributed to the presence of the parasite in the kidney tissues of participants in this study probably due to latent infection. These results have a number of similarities with some studies investigated the prevalence of T. gondii in patients with renal failure [38,39]. According to our knowledge, no data were available regarding detection of T. gondii using urine samples in Saudi Arabia, therefore, these findings may be considered for confirmation of latent toxoplasmosis infection especially when low antibody response in patients reduce the sensitivity of serologic tests.
Limitation(s)
It was plausible that a number of limitations could have influenced the result obtained, first was the low sample size as the study was self-funded and due to the limited facilities only 22 samples were considered out of 74 IgG anti-T gondii positive cases [19]. The second was the use of only IgG positive samples, it could be more sound to use IgM positives as well as antigen positive and isolated parasite samples to ensure the existence of DNA. General limitations of molecular work like shortages of reagents due to high cost are considered.
Conclusion(s)
The study concluded that LAMP assay based on B1 and SAG2 gene had a greater sensitivity for T. gondii detection. On the other hand, results propose that the LAMP detection of T. gondii DNA in urine may be an appropriate method for the diagnosis of toxoplasmosis in chronic infection rather than early infection. Authors recommended that a specialised laboratory for toxoplasmosis should be established in the area, in order to undertake advanced research and diagnostic services.