Head and neck cancer is the second most common cancers in the Indian population with 77,000 cases were diagnosed per year [1]. It can disfigure the face, strip away the voice and rob one of his/her basic abilities to eat, drink, swallow and the psychosocial impact can be extremely depressing. Epidemiological studies revealed that HNC occurs through a complex process involving exposure to carcinogens from smoking and alcohol consumption [2-4]. Tobacco and alcohol consumption are the major risk factors for HNC likely due to DNA damaging process [5]. Cigarette contains some amounts of chemicals containing many known carcinogens; carcinogenic role of cigarette varies depending upon the cigarette product [6]. The chemical products of alcohol metabolism are also explored to be toxic and hypothesised to cause DNA damage that lead to cancers [7].
It is hypothesised that susceptibility to disease development is correlated with efficiency of DNA repair and cell cycle control or a combination of both of these mechanisms. Individuals thus differ in their capacity to repair DNA damage on exposure to carcinogens [8]. Some selected DNA repair enzymes play a role in HNC.
Genetic polymorphisms in some DNA repair proteins are associated with number of malignancies including HNC [20]. Hence, the present study aimed to draw an association (if any) between polymorphic 347 bp and 445 bp variants of XRCC1 gene and the risk of HNC in a Southern Indian population.
Materials and Methods
Study Subjects and Ethical Approval
The present case control study was conducted at the Department of Radiotherapy, Government General Hospital, Guntur, India, study period was of two years; from June 2012 to June 2014. A total of 76 males and 44 female newly diagnosed HNC cases (newly diagnosed with Stage I and II) were selected by using simple random sampling. Data obtained regarding attitudes and habits considered for risk assessment and XRCC1 gene SNP were analysed in 30 HNC cases and 30 healthy subjects (controls) to compare the frequency of polymorph variants. Peripheral blood obtained from both cases and controls. During examination, the cases and controls did not receive any medicaments like antibiotics or steroids.
The inclusion criterion for the cases was the presence of histopathologically diagnosed head and neck squamous cell carcinoma of the head and neck cancers who were in Stage I and II without any other diseases. Only 30-60-year-old patients were enrolled for the study. The inclusion criterion for the controls was absence of prior history of cancer. Stage III and Stage IV HNC cases and cases with other disease like Diabetes, heart problems etc., were excluded from the study. Healthy controls were recruited from members of different community centers. The Institutional Ethics Committee approved the study and written consent was obtained from each patient and control group before enrolling into the study.
SNP Detection by Tetra ARMS PCR
DNA was isolated by chloroform extraction after overnight incubation with proteinase K at 37°C. A protocol of HiPurA™ (MB504) Blood Genomic DNA Miniprep Purification Kit was used for extraction of DNA [21].
Estimation of DNA
The isolated DNA was studied for quantity and quality analysis before using it for molecular analyses. The estimation of DNA was done with the spectrophotometer. Cuvette was filled with TE buffer, Zero the spectrophotometer at 260 nm with this blank. 1 microlitre DNA sample were required and usually diluted before measuring its absorbance. Diluting the DNA sample 1μL: 999 μL of TE buffer; mixed the dilution thoroughly. Measured the Optical Density (OD), a ratio of the OD-260 nm/OD-280 nm is an indicator of DNA purity. A ratio of 1.8 or higher indicates minimal protein contamination [22], multiplied the resulting OD by 50 μL/mL. For a 1: 1000 dilution, the mass of DNA was equal to 1 μL. Similarly, the same sample was measured at 280 nm.
Agarose Gel Electrophoresis
Agarose, 50X TAE buffer, 6X gel loading buffer and Ethidium bromide were used for Agarose Gel Electrophoresis.
Tetra Primer-amplification Refractory Mutation System (ARMS)-PCR Method
Tetra primer ARMS PCR works with two pairs of primers to amplify two alleles in one PCR reaction as described on the basis of previous study [23].
Statistical Analysis
Deviations from the Hard-Weinberg Equilibrium (HWE) were tested by using online calculator, chi-square test was used to compare between two variables. The odds ratio with corresponding 95% Confidence Interval (CI) was used to examine the association between XRCC1 polymorphism and HNC risk. For all statistical analyses performed in present thesis, p-values ≤0.05 were considered significant.
Results
In the present study, the attitudes and habits considered for risk assessment in 120 (76 male and 44 female) HNC Cases [Table/Fig-1]. Among 120 cases 37 (48.68%) males and 22 (50%) females were in the age group of 30-45 years of age, and the rest i.e., 39 (51.31%) males and 22 (50%) female cases were in the age group of 46-60 years. It was also observed that the percentage of malignancy was higher in the age group of 45-60 years than in the 30-45 year’s age group. Out of 76 males 12 (15.78%) and out of 44 females 5 (11.36%) cases had a family history of malignancy. Out of 120 HNC cases, only 17 cases (14.16%) had a family history of HNC and other 103 cases (85.83%) had no family history of HNC. Thus, the incidence of malignancy appears to be not associated with the family history.
Out of 120 HNC cases 37 (84.09%) female cases were non-smokers. Out of 120 HNC cases 72 (60%) cases have the habit of smoking. Among male cases alone, 75.50% constitute smokers. A total of 57 cases, consisting of 18 (23.68%) males and majority of females (39; 88.63%) responded that they never consume alcohol.
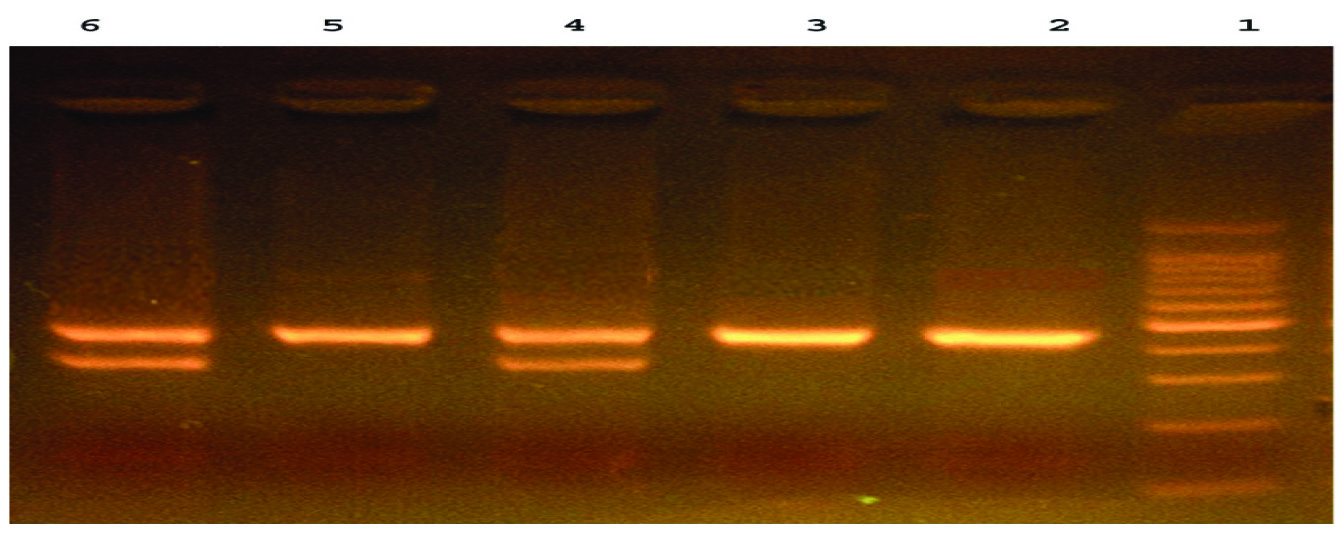
The gel picture shows the DNA fragments representative for Homozygote Mutant G/G (445bp and Heterozygote A/G (347bp/445bp) respectively [Table/Fig-2]. Genotype distribution of Arg194Trp (χ2=3.380; p=0.064) and Arg399Gln (χ2=0.188; p=0.713) polymorphisms corresponds to Hard-Weinberg Equilibrium.
Characteristics of 120 (76 male and 44 female) head and neck cancer cases.
| Characteristics | Males 76 (63.33%) | Females 44 (36.66%) |
|---|
| Age (years) | 30-45 | 37 (48.68) | 22 (50) |
| 45-60 | 39 (51.31) | 22 (50) |
| Smoking | Non-smoker | 11 (14.47) | 37 (84.09) |
| 1-5 Cigarettes/day | 23 (30.26) | 7 (15.90) |
| 5-10 Cigarettes/day | 27 (35.52) | 0 |
| >10 Cigarettes/day | 15 (19.73) | 0 |
| Alcohol drinking | Never/ occasionally | 18 (23.68) | 39 (88.63) |
| 1-2 pegs/day | 21 (27.63) | 5 (11.36) |
| 3-7 pegs/day | 16 (21.05) | 0 |
| >7 pegs/day | 21 (27.63) | 0 |
A representative agarose gel picture of digested PCR products. 100bp marker; Lane 1: Ladder; Lane 2, 3,5: Homozygote Mutant G/G (445 bp); Lane 4, 6 Heterozygote A/G (347 bp/445 bp).
The XRCC1 SNP sequence information and designed primers for tetra primer ARMS-PCR reactions are shown in the [Table/Fig-3]. The XRCC1 gene SNPs were analysed in 30 healthy subjects (controls) and 30 HNC cases, randomly selected. The frequency homozygous genotypes (AA and GG) and allele frequencies and odds ratios are presented in [Table/Fig-3]. The heterozygous genotype (AG) representing single nucleotide polymorphism was observed in 80% controls and 63.33% in HNC cases. The χ2 value calculated for heterozygotes frequency in controls and cases did not differ significantly showing that there was no direct association between SNPs of XRCC1 gene and outcome of HNC. The frequency of heterozygous alleles (χ2 value and p-value at ≤0.05) also revealed no significant positive association between SNPs of XRCC1 and HNC [Table/Fig-4].
XRCC1 SNP sequence information and designed primers for tetra primer ARMS-PCR reactions (>gnl | dbSNP | rs25487 | allelePos=501 | total Len=1001 | taxid=9606 | snpclass=1 | alleles=’A/G’ |mol=Genomic|build=137)
| XRCC1 SNP Sequence Information |
|---|
| Forward inner primer (A allele) | GTCGGCGGCTGCCCTCACA |
| Reverse inner primer (G allele) | GGCGTGTGAGGCCTTACCGCC |
| Forward outer primer (5’-3’) | CCCTGCGCCGCTGCAGTTTC |
| Reverse outer primer (5’-3’) | CGTGAGCCACTGCACCGGGA |
Single nucleotide (347 bp and 445 bp) polymorphism of XRCC1 gene in HNC cases and controls.
| Genotypes | Cases(n=30) | Controls(n=30) | Odds ratio andClass Interval (CI) | χ2-valuep-value |
|---|
| A A | 0 | 0 | 0.4318CI (0.135, 1.381) | 2.052(0.152) |
| A G(Heterozygous) | 19 (63.33%) | 24 (80%) |
| G G(Homozygous) | 11 (36.66%) | 6 (20%) |
| Allele frequency | n=60 | 0.695CI (0.217, 2.223) | 0.906(0.345) |
| A | 19 (31.66%) | 24 (40) |
| G | 41 (68.33%) | 36 (60) |
| Hard-Weinberg Equilibrium (HWE) | 0.011 | 0.0003 |
Statistically significant difference at (p≤0.05).
Discussion
Out of 120 HNCs, 72(60%) were smokers. Tobacco smoke contains over 2,000 chemicals; many of them are directly related to the cause of cancer. Mostly cigarettes are kept between lips and teeth from where chemicals are gradually absorbed after dilution with saliva. So, particularly the side of the tongue, the floor of the mouth and alveolus are the sites of maximum result, and thus are maximally affected. One of the previous studies [17], stated that 64% of cancer cases of their study group are smokers and in another study, [24] stated that 80% of their cancer patients are reported as tobacco users and 75.7% consume alcohol respectively.
In the present study, out of 120 HNC Cases 63 cases (52.5%) consumed alcohol. Alcohol consumption along with tobacco increases the risk of cancers of the HNCs and enhanced effect more than the independent effect of either drinking or/and smoking [25].
DNA repair mechanisms play an important role in the protection of cells from DNA damage and maintenance of genome integrity. The genetic polymorphism of some DNA repair genes is associated with malignancies including HNC. Khlifi R et al., reported the link between variations such as single nucleotide polymorphisms of DNA repair genes (XRCC1, ERCC2 and XRCC3) and increased risk of HNC. The inferences drawn from the SNP’s of DNA repair genes (XRCC1, ERCC2, XRCC3) vary considerably [20].
The human’s XRCC1 gene is located at the long arm of 19 chromosome and it encodes a 633-amino acid long protein. This functions as scaffolding protein for other proteins [26]. The two polymorphisms C→A substitution (Arg 194 Trp) and G→A substitution (Arg 399 Gln) of XRCC1 were investigated in several studies [26,27].
Sturgis EM et al., found an elevated risk for less common genotype (Gln, Gly vs Arg/Gln or Arg/Arg) in Texas population [14]. Olshan AF et al., observed a decreased risk of HNC for homozygous genotype (Gln/Gln) among White Americans [28]. Recently a statistically significant association was found for XRCC1 codons 399, 194 variants and HNC American (New York) population. No significant difference was found between patients and controls with respect to polymorphism of XRCC1 (Arg 194 Trp and Arg 399 Gln) in a Hungarian population [26].
Lou Y et al., found significant differences between study heterogeneities in the Asians and their insignificance in Caucasians [29]. In the present study, also the frequency of heterozygous genotype (A/G) of XRCC1 gene in both controls and HNC cases showed no significant difference and hence, no association between them. Based on the present study results, we support the Khlifi R et al., statement that the association between XRCC1 gene polymorphisms and HNC cases is still confusing, despite of large number of studies in well-characterised populations [20].
Limitation
Present study limited by a smaller sample size, which might result in a lack of ability to detect the possible risk for XRCC1 polymorphism in HNC. The present study was based on unadjusted estimates, while a more exact analysis could be performed if individual data were available.
Conclusion
The present study confirms the principle role of cigarette smoking and alcohol consumption in HNC. There is a significant and positive interaction between these two factors. The results of the current study suggest that 347 bp and 445 bp variants of XRCC1 gene polymorphism are not involved in HNC susceptibility. In addition, further studies are required for evaluating the effect of gene to gene and gene to environment interactions on these gene polymorphisms with HNC susceptibility in Indian population.
Statistically significant difference at (p≤0.05).