Association of Lin-28A rs3811464 Variant with Susceptibility to Type 2 Diabetes
Mona Khodabandeh1, Hamid Ghaedi2, Behnam Alipoor3, Taghi Golmohammadi4
1 Postgraduate Student, Department of Cell and Molecular Biology, Faculty of Biological Sciences, Kharazmi University, Tehran, Iran.
2 Assistant Professor, Department of Medical Genetics, Faculty of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran.
3 Assistant Professor, Department of Laboratory Sciences, Faculty of Paramedicine, Yasuj University of Medical Sciences, Yasuj, Iran.
4 Associate Professor, Department of Biochemistry, School of Medicine, Tehran University of Medical Sciences, Tehran, Iran.
NAME, ADDRESS, E-MAIL ID OF THE CORRESPONDING AUTHOR: Dr. Taghi Golmohammadi, Associate Professor, Department of Biochemistry, Faculty of Medicine, School of Medicine, Tehran University of Medical Sciences, Tehran-1417653761, Iran.
E-mail: golmoham@sina.tums.ac.ir
Introduction
It has been suggested that Lin-28A and the let-7 microRNA family (Lin-28/let-7 axis) play a critical role in the control of glucose metabolism, insulin sensitivity and resistance to diabetes.
Aim
This case-control study aimed at evaluating the association between Lin-28 rs3811464 polymorphism and the susceptibility to Type 2 Diabetes (T2D) in a sample of Iranian population.
Materials and Methods
This study involved 172 T2D patients and 160 non-diabetic age and gender-matched controls. Lin 28A rs3811464 genotypes were determined by Polymerase Chain Reaction–Restriction Fragment Length Polymorphism (PCR-RFLP) technique.
Results
The results showed that the frequency of the AA genotype was significantly higher in control subjects than in diabetic patients (13.12% vs. 4.65%). In addition, binary logistic regression analysis revealed that rs3811464-AA genotype was significantly associated to T2D after adjustment for BMI, age and lipid profiles. Indeed, subjects with AA genotype were less likely to develop T2D than GG and AG subjects (OR of 0.26, 95% CI 0.10-0.66, p=0.005).
Conclusion
The findings of our study suggest that the Lin 28A rs3811464 is associated with type 2 diabetes susceptibility and subjects with AA genotypes were less likely to develop T2D diabetes.
Diabetes risk, Genetic association study, Single nucleotide polymorphism
Introduction
Diabetes mellitus is a major global health problem which imposes a huge economic burden on societies [1]. It has been shown that in 2013, 382 million people had diabetes and it is expected to rise to 592 million by 2035 [2]. Type 2 Diabetes (T2D) is the most common form of diabetes which is characterized by chronic hyperglycaemia due to defects in insulin secretion or its action [3,4]. As a metabolic disorder, T2D leads to various complications such as cardiovascular disorders, retinopathy and nephropathy and thereby causes high rate of morbidity, disability and mortality worldwide [5,6]. T2D is a multifactorial disease resulting from a complex interaction between genetic and environmental factors [6]. Many studies identified different loci implicated in the genetic predisposition to T2D, but the genetics of T2D remained to be elusive and not well understood [7,8].
Lin-28 (including Lin-28A and Lin-28B) is an evolutionarily conserved RNA-binding protein that plays key roles in multiple cellular developmental processes [9,10]. New studies suggested that Lin-28A and its homolog Lin-28B and the let-7 microRNA family (Lin-28/let-7 axis) play a direct role in the regulation of glucose metabolism [9,11,12]. Zhu H et al., in a study showed that Lin-28A in transgenic mouse model exhibited enhanced glucose uptake in peripheral tissues. Indeed their findings revealed that Lin-28A overexpression results in insulin sensitivity, enhanced glucose tolerance, and resistance to diabetes [11]. Moreover, conditional deletion of Lin-28A in skeletal muscles lead to insulin resistance and impaired glucose uptake, suggesting that Lin-28 is physiologically required for normal glucose homeostasis [12]. Zhu H et al., also showed that in contrast to the phenotypes seen in Lin-28A transgenic mouse model, inducible let-7 transgenic mice have hyperglycaemia and glucose intolerance. These findings imply that Lin-28 may increase glucose uptake by suppressing let-7 [12].
It has been suggested that Single Nucleotide Polymorphisms (SNPs) in let-7/Lin-28 axis could affect the interaction of let-7 and Lin-28 and result in more susceptibility to diseases [13-16]. Chen AX et al., reported that rs3811463 polymorphism could lead to differential regulation of Lin-28 by let-7 and have a significant effect on the risk of breast cancer. They showed that the C allele of rs3811463 weakened let-7–induced repression of Lin-28 mRNA, resulting in increased production of Lin-28 protein [13]. However, they did not found a significant association between rs3811464 and breast cancer in a sample of Chinese population. The rs3811464 is located in the promoter region of Lin-28 that is 126 bp upstream of the transcriptional start site [13]. Till date only one study has investigated the association of this polymorphism with T2D susceptibility which failed to find differences in the frequencies of rs3811464 genotypes between diabetic patients and control subjects [17].
Considering the important role of the Lin-28 in the insulin and glucose metabolism and also the impact of SNPs on the Lin-28 expression and function, in this case-control study we aimed to investigate the possible association between the Lin-28A rs3811464 polymorphism with the susceptibility to T2D in a sample of Iranian population.
Materials and Methods
This case-control study was conducted from March 2016 to November 2016 and involved 172 T2D patients and 160 non-diabetic age and gender-matched controls. All subjects were recruited from Shahid Taleghani hospital (Tehran, Iran). Diabetes is defined as Fasting Blood Sugar (FBS) ≥126 mg/dL and two hours glucose ≥200 mg/dL. The control group was chosen among the healthy subjects without a family history of diabetes who had FBS level of <100 mg/dL and Haemoglobin A1c (HbA1C) <5.7%. Individuals with type 1 diabetes mellitus, cardiovascular disorders, renal and hepatic diseases and patients with any malignancies were excluded from the study. The study was approved by the Ethical Committee of Shahid Beheshti University of Medical Sciences and written informed consent was obtained from all subjects before enrollment in the study. Demographic characteristics of study population were collected by interviewer-administered questionnaire and the medical records.
Biochemical Measurements
Total cholesterol, triglyceride, HDL–cholesterol, and FBS levels were measured using the routine laboratory methods. LDL-cholesterol was calculated using the Friedewald equation. Moreover, HbA1C level was evaluated by high performance liquid chromatography method.
DNA Extraction and Genotyping
Genomic DNA was isolated from the whole blood samples by salting out method.
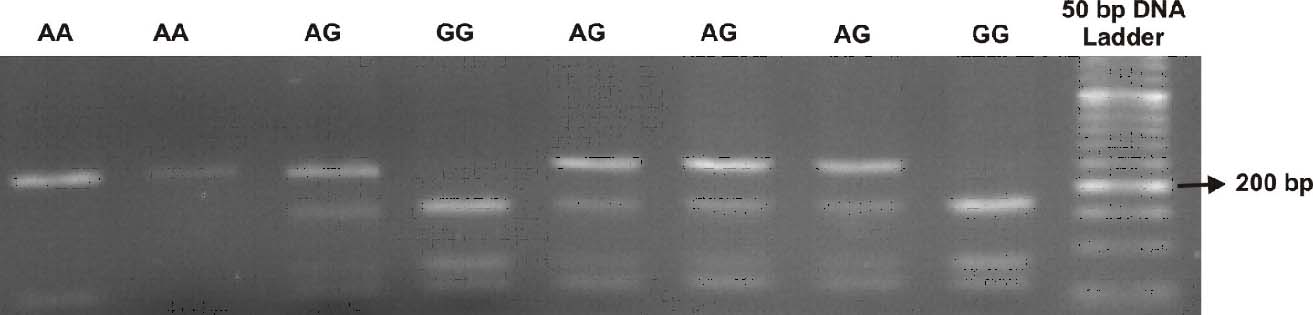
Lin-28A rs3811464 genotypes were determined by PCR-RFLP technique. PCR amplification was carried out using the following primers: forward 5’- AGGCAAAGGGTTGGTTCGG -3’ and reverse 5’- CACCTGTATCTGCTTTGGGGAC -3’. The thermal cycling conditions were as follows: 94°C for 5 minutes followed by 30 cycles comprising of 95°C for 40 seconds, annealing time at 61°C for 40 seconds and extension at 72°C for 40 seconds with a final extension time of seven minutes at 72°C. The PCR product (368bp) was digested with 10 units of NlaIV restriction enzyme and then the fragments were separated on a SYBR Green stained 3% agarose gel. The A allele produced 264 bp and the G allele yielded 175 and 89 bp fragments after digestion. While the heterozygous AG was characterized by three bands of 264, 175 and 89 bp [Table/Fig-1]. All genotypes produced 64, 21 and 19 bp bands after digestion. Fragments smaller than 50 bp were not visualized [Table/Fig-1].
PCR-RFLP based genotypes of Lin-28A rs3811464 variant.
Bioinformatic Analysis
For further investigation, the rs3811464 variant was analyzed in the genomics context. A Multiple Sequence Alignment (MSA) performed to assess the sequence conservation [18]. Moreover, the haplotype blocks and SNPs in strong Linkage Disequilibrium (LD) were obtained from HaploReg v4 [19]. To more dissect the effect of rs3811464 different alleles on Lin-28-3’UTR secondary structure, the Srna software was used to calculate partition function for sampling 1000 Boltzmann ensemble structures and clustering of Lin-28-3’UTR secondary structure with alternative and wild-type alleles [20].
Statistical Analysis
All statistical analysis was done by Statistical Software Package (SPSS) version 18.0. The continuous and categorical variables were expressed as mean ± SD and percentages (n), respectively. Student’s t-test was used to compare the quantitative variables. Compatibility of genotype frequencies with Hardy–Weinberg equilibrium expectations and comparisons of the categorical variables were evaluated by chi-square test. Independent Odds Ratio (OR) and its 95% confidence interval (CI) were calculated using unconditional binary logistic regression after adjusting for the confounders of age, BMI and lipid levels. A p-value <0.05 were considered to be statistically significant.
Results
Characteristics of the Study Population
Clinical and biochemical characteristics of total 332 subjects are summarized in [Table/Fig-2].
Clinical characteristic of the T2D patients and control subjects.
| Parameter | Control subjects(n = 160) | T2D(n = 172) | p-value |
|---|
| Age (years) | 53.06±6.62 | 54.46±8.81 | 0.103 |
| Sex (Male/Female) | 78/82 | 90/82 | 0.58 |
| BMI (kg/m2) | 27.56±3.93 | 29.26±4.37 | <0.001 |
| Systolic blood pressure (mmHg) | 124.05±17.96 | 128.76±19.73 | 0.023 |
| Diastolic blood pressure (mmHg) | 75.88±11.34 | 79.44±15.33 | 0.017 |
| Fasting plasma glucose (mg/dL) | 86.49±8.76 | 140.51±35.47 | <0.001 |
| HbA1C% | 5.36±0.37 | 7.62±1.04 | <0.001 |
| Total cholesterol (mg/dL) | 160.50±44.87 | 168.30±38.92 | 0.090 |
| Triglyceride (mg/dL) | 138.35±71.08 | 158.91±76.34 | 0.011 |
| HDL-C (mg/dL) | 43.87±10.20 | 41.55±9.36 | 0.031 |
| LDL-C (mg/dL) | 87.59±21.58 | 88.36±26.93 | 0.774 |
There were no significant differences between groups for age and sex. T2D patients had significantly higher values for BMI (p<0.001), systolic blood pressure (p=0.023), diastolic blood pressure (p=0.017), triglycerides (p=0.011), FBS (p<0.001), HbA1C (p<0.001) and lower level of HDL (p=0.031) than the control subjects. Moreover, there was no statistically significant difference between T2D and normal subjects for LDL (p=0.774) and total cholesterol levels (p=0.09).
Association of rs3811464 variant with susceptibility to T2D: genotype distribution and allele frequency of the rs3811464 variant in T2D patients and control subjects are shown in [Table/Fig-3]. The genotype distributions in the study population was compatible with Hardy–Weinberg equilibrium (p>0.05).
Genotype distribution and allele frequency rs3811464 polymorphism.
| Genotypes | Control subjects(n=160) | T2D(n=172) | p-value | OR (95% CI) |
|---|
| Co-dominant |
| GG | 55 (34.37%) | 64 (37.2%) | 0.024 | |
| AG | 84 (52.5%) | 100 (58.13%) | |
| AA | 21 (13.12%) | 8 (4.6%) | |
| Dominant model |
| AG+AA | 105 (65.62%) | 108 (62.79%) | 0.59 | 0.88 (0.56-1.38) |
| GG | 55 (34.38%) | 64 (37.21%) |
| Recessive model |
| GG+AG | 139 (86.88) | 164 (95.35%) | 0.006 | 0.32 (0.13-0.75) |
| AA | 21 (13.12%) | 8 (4.65%) |
| Over-dominant model |
| GG+AA | 76 (47.5) | 72 (41.86%) | 0.30 | 0.79 (0.51-1.22) |
| AG | 84 (52.5%) | 100 (58.14%) |
| Allele |
| G | 194 (60.63%) | 228 (66.27%) | 0.14 | 1.27 (0.93-1.75) |
| A | 126 (39.37%) | 116 (33.73% |
Although the A allele was more observed among the control subjects, rs3811464 allele distribution was not significant between the groups (p=0.14). The genotype frequencies of the rs3811464 polymorphism was differently distributed between T2D and control subjects (p=0.02). Our results showed that the frequency of the AA genotype was significantly higher in control subjects than in T2D patients (13.12% vs. 4.65%). In addition, dominant, recessive and over-dominant models of inheritance were evaluated. The results showed that the rs2910164 variant was protective against T2D under recessive model [Table/Fig-3]. Binary logistic regression analysis was done to evaluate the independent association of rs3811464 variant with T2D. Our analysis revealed that rs3811464-AA genotype was significantly associated to T2D after adjustment for BMI, age and lipid profiles [Table/Fig-4]. Indeed, subjects with AA genotype were less likely to develop T2D than GG and AG subjects (OR of 0.26, 95% CI 0.10-0.66, p=0.005).
Logistic regression analysis of variables associated with the risk of T2D.
| Parameter | p-value | OR | 95% CI |
|---|
| Age | 0.077 | 1.027 | (0.997-1.05) |
| BMI | 0.001 | 1.100 | (1.038-1.16) |
| Triglyceride | 0.127 | 1.003 | (0.999-1.00) |
| Total cholesterol | 0.161 | 1.005 | (0.998-1.01) |
| LDL-C | 0.267 | 0.993 | (0.982-1.00) |
| rs3811464-AA | 0.005 | 0.26 | (0.10-0.66) |
BMI: body mass index; LDL-C: Low-density lipoprotein cholesterol; CI: confidence interval; OR: Odds ratio. Type 2 diabetes was considered as dependent variable.
Bioinformatic Analyses Results
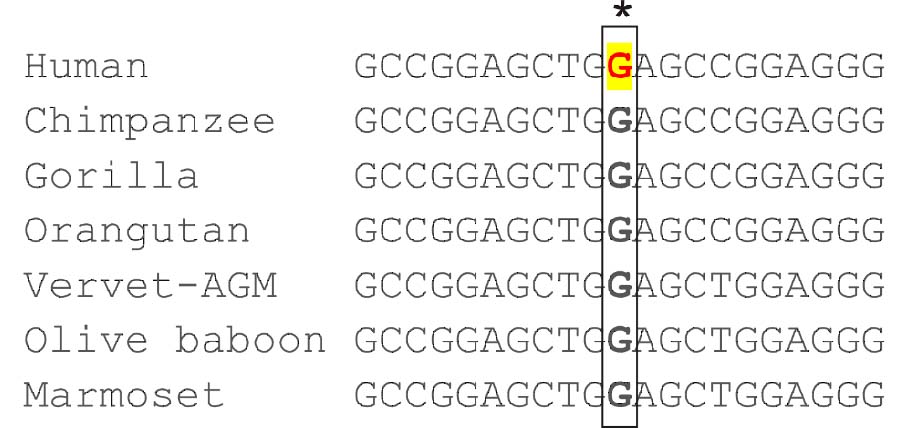
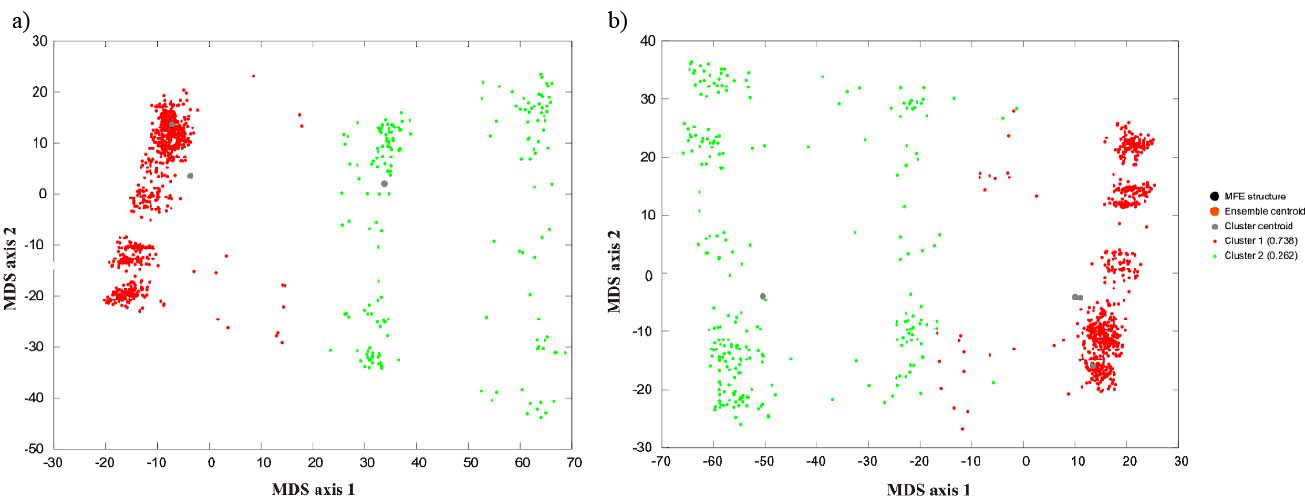
MSA analysis showed that rs3811464 variant is well conserved among the selected primates [Table/Fig-5]. The HaploReg analyses revealed several correlated polymorphism in strong LD (r2>0.9) in Asian (ASN) and European (EUR) population from the 1000 Genomes Project with rs2910164 variant [Table/Fig-6]. We draw 1000 samples from the ensemble of secondary structures of Lin-28-3’UTR in proportion to Boltzmann weights for the A and the G alleles of rs3811464. As illustrated by Multi Dimensional Scaling (MSD) plots, the 3’UTR conform two distinct clusters with A allele and four distinct clusters in the case of G allele [Table/Fig-7]. This result indicates the structural effects of rs2910164 on the host gene secondary structure.
Multiple sequence alignment for Lin-28A rs3811464 variant and flanking sequence. The asterisks denote conserved nucleotides.
Polymorphisms linked to rs3811464 variant in HaploReg.
| Variant | LD(r2) | Proteins bound | Motifs changed | GENCODE genes |
|---|
| rs12744785 | 0.91 | - | Ets,Mrg,Tgif1 | 12kb 5’ of LIN-28A |
| rs12402357 | 0.91 | - | AhR::Arnt,Arnt | 11kb 5’ of LIN-28A |
| rs11247948 | 0.94 | - | 4 altered motifs | 11kb 5’ of LIN-28A |
| rs11247949 | 0.96 | - | 11 altered motifs | 11kb 5’ of LIN-28A |
| rs7556500 | 0.97 | - | 4 altered motifs | 7.7kb 5’ of LIN-28A |
| rs11577535 | 0.95 | GATA3 | 21 altered motifs | 6.1kb 5’ of LIN-28A |
| rs7513273 | 0.95 | FOXA1 | 10 altered motifs | 5.9kb 5’ of LIN-28A |
| rs7517877 | 0.92 | CJUN | 5 altered motifs | 5.5kb 5’ of LIN-28A |
| rs7549684 | 0.94 | CJUN | - | 5.5kb 5’ of LIN-28A |
| rs7552060 | 0.95 | CJUN | - | 5.3kb 5’ of LIN-28A |
| rs11247952 | 0.94 | - | Nkx2, Nkx3, SETDB1 | 5.2kb 5’ of LIN-28A |
| rs11247953 | 0.95 | - | 18 altered motifs | 5kb 5’ of LIN-28A |
| rs6669670 | 0.95 | - | Mtf1 | 3.7kb 5’ of LIN-28A |
| rs12759853 | 0.94 | - | 4 altered motifs | 3.6kb 5’ of LIN-28A |
| rs35347695 | 0.93 | - | - | 2.4kb 5’ of LIN-28A |
| rs7530114 | 0.98 | BAF170, BRG1, NRSF | Ik-1,ZBTB7A | 2.2kb 5’ of LIN-28A |
| rs12747426 | 0.96 | - | BDP1,HDAC2 | 1.6kb 5’ of LIN-28A |
| rs12122703 | 0.96 | MAX | 9 altered motifs | 1.5kb 5’ of LIN-28A |
| rs3811464 | 1 | - | 5 altered motifs | 125bp 5’ of LIN-28A |
The secondary structures of Lin-28-3’UTR. a,b) depicts the Lin-28-3’UTR centroid structures and MSD plots of the structural ensemble with A and G allele, respectively.
Discussion
In our knowledge, this is the first case-control study which evaluates the association between Lin-28A rs3811464 polymorphism and the susceptibility to T2D in a sample of Iranian population. The finding of our study shows that the AA genotype frequency was significantly higher in control subjects than in T2D patients. Indeed, our results emphasize that subjects with AA genotype were less likely to develop T2D than GG and AG subjects. This finding is not in agreement with the results of Zhang J et al., study in the Han population which failed to find significant differences in the frequencies of rs3811464 genotypes between diabetic patients and control subjects [17]. One possible reason for such inconsistency can be explained by the genetic differences between Iranian and China populations and also in the study sample size.
Emerging evidence has reported that Lin-28 plays a critical role in the control of glucose metabolism, insulin sensitivity and resistance to diabetes. Zhu H et al., reported that the Lin-28A transgenic mice showed enhanced glucose uptake and lower fed state glucose. In addition they revealed that, shRNA knockdown of Lin-28A in C2C12 led to a reduction in labeled glucose uptake [11]. Similarly, Zhu H et al., in another study showed that Lin-28A transgenic mice cleared glucose more efficiently during glucose and insulin tolerance testing. In addition, it has been shown that glucose hemostasis would be impaired in mice with muscle-specific Lin-28A knock-out and inducible let-7 [12]. Taken together, their results demonstrated that Lin-28 may increases glucose uptake by suppressing let-7. Shinoda G et al., reported that Lin-28A and Lin-28-B deficiency in knockout mice led to growth defects and aberrations in glucose metabolism [21].
Due to the important role of the Lin-28/let-7 axis in the regulation of glucose metabolism, genetic variation in Lin-28 could affect the interaction of let-7 and Lin-28 and result in more susceptibility to diseases. For example, the allelic variants of rs3811463 could alter the local mRNA secondary structure including that of the let-7 binding site [13]. Chen AX et al., identified a SNP, rs3811464, in the promoter region of Lin-28 that was 126 bp upstream of the transcriptional start site of Lin-28 [13]. In contrast with the results of Zhang J et al., our results emphasize that subjects with rs3811464-AA genotype were less likely to develop T2D [17]. This polymorphism was not located near the binding sites of miRNAs targeting Lin-28 such as Let-7. Therefore, other regulatory mechanisms may contribute to the impact of Lin-28 rs3811464 on the regulation of let-7/Lin-28 loop and development of T2D disease. For example, because the rs3811464 is located in the 59 flanking region (promoter) of the Lin-28 gene, allele of this variation could lead to alternation of the transcriptional factor binding site. Further functional studies will be necessary to obtain a deeper comprehension about rs3811464 association with diabetes.
Our bioinformatics analyses also provide more evidence on the association of rs3811464 variant with T2D susceptibility. MSA analysis showed that the rs3811464 variant and flanking sequence is conserved across primate taxa. Study reported that SNPs within conserved DNA regions could pursue phenotypic consequences [22]. In addition, it is widely accepted that region-based analysis has several advantages than single-SNP analysis to identify disease-associated loci [23]. Our result identified several correlated polymorphism in strong LD with the rs2910164 polymorphism. Further investigations into the LD relationships between different variants in different parts of the genome could enhance the ability to identify association with a disease phenotype [24]. Moreover, modeling secondary structure of Lin-28-3’UTR in the presence of rs3811464 different allele revealed that, changing from A to G changed the energy of secondary structures markedly. Such SNP-mediated changes in secondary structure may elicit functional consequences.
Limitation
As a limitation our study suffers from limited sample size, multicenter and additional studies with a larger sample size will be necessary to confirm the association of these variant with T2D susceptibility.
Conclusion
The results of our study suggest that the Lin-28A rs3811464 is associated with T2D susceptibility and subjects with AA genotype were less likely to develop T2D.
[1]. Dall TM, Yang W, Halder P, Pang B, Massoudi M, Wintfeld N, The economic burden of elevated blood glucose levels in 2012: diagnosed and undiagnosed diabetes, gestational diabetes mellitus, and prediabetesDiabetes Care 2014 37(12):3172-79. [Google Scholar]
[2]. Guariguata L, Whiting D, Hambleton I, Beagley J, Linnenkamp U, Shaw J, Global estimates of diabetes prevalence for 2013 and projections for 2035Diabetes Research And Clinical Practice 2014 103(2):137-49. [Google Scholar]
[3]. Khodabandehloo H, Gorgani-Firuzjaee S, Panahi G, Meshkani R, Molecular and cellular mechanisms linking inflammation to insulin resistance and β-cell dysfunctionTranslational Research 2016 167(1):228-56. [Google Scholar]
[4]. Tripathy D, Chavez AO, Defects in insulin secretion and action in the pathogenesis of type 2 diabetes mellitusCurrent Diabetes Reports 2010 10(3):184-91. [Google Scholar]
[5]. McClelland AD, Kantharidis P, MicroRNA in the development of diabetic complicationsClinical Science 2014 126(2):95-110. [Google Scholar]
[6]. Ahlqvist E, Van Zuydam NR, Groop LC, McCarthy MI, The genetics of diabetic complicationsNature Reviews Nephrology 2015 11(5):277-87. [Google Scholar]
[7]. Alipoor B, Meshkani R, Ghaedi H, Sharifi Z, Panahi G, Golmohammadi T, Association of miR-146a rs2910164 and miR-149 rs2292832 Variants with Susceptibility to Type 2 DiabetesClinical Laboratory 2016 62(8):1553-61. [Google Scholar]
[8]. Ghaedi H, Tabasinezhad M, Alipoor B, Shokri F, Movafagh A, Mirfakhraie R, The pre-mir-27a variant rs895819 may contribute to type 2 diabetes mellitus susceptibility in an Iranian cohortJournal Of Endocrinological Investigation 2016 39(10):1187-93. [Google Scholar]
[9]. Nguyen LH, Zhu H, Lin28 and let-7 in cell metabolism and cancerTranslational Paediatrics 2015 4(1):4 [Google Scholar]
[10]. Jiang S, Baltimore D, RNA-binding protein Lin28 in cancer and immunityCancer Letters 2016 375(1):108-13. [Google Scholar]
[11]. Zhu H, Shah S, Shyh-Chang N, Shinoda G, Einhorn WS, Viswanathan SR, Lin28a transgenic mice manifest size and puberty phenotypes identified in human genetic association studiesNature Genetics 2010 42(7):626-30. [Google Scholar]
[12]. Zhu H, Shyh-Chang N, Segrè AV, Shinoda G, Shah SP, Einhorn WS, The Lin28/let-7 axis regulates glucose metabolismCell 2011 147(1):81-94. [Google Scholar]
[13]. Chen AX, Yu KD, Fan L, Li JY, Yang C, Huang AJ, Germline genetic variants disturbing the Let-7/LIN28 double-negative feedback loop alter breast cancer susceptibilityPLoS Genet 2011 7(9):e1002259 [Google Scholar]
[14]. Zhang Y, Wang R, Miao L, Jiang H, Yuan H, Ma H, Genetic variants in let-7/Lin28 modulate the risk of oral cavity cancer in a Chinese Han populationScientific Reports 2014 :4 [Google Scholar]
[15]. Wen J, Liu H, Wang Q, Liu Z, Li Y, Xiong H, Genetic variants of the LIN28B gene predict severe radiation pneumonitis in patients with non-small cell lung cancer treated with definitive radiation therapyEuropean Journal of Cancer 2014 50(10):1706-16. [Google Scholar]
[16]. Ye Y, Madison B, Wu X, Rustgi AK, A LIN28B polymorphism predicts for colon cancer survivalCancer Biology & Therapy 2012 13(14):1390-95. [Google Scholar]
[17]. Zhang J, Zhang L, Fan R, Guo N, Xiong C, Wang L, The polymorphism in the let-7 targeted region of the Lin28 gene is associated with increased risk of type 2 diabetes mellitusMolecular and Cellular Endocrinology 2013 375(1):53-57. [Google Scholar]
[18]. Sievers F, Higgins DG, Clustal Omega, accurate alignment of very large numbers of sequencesMultiple Sequence Alignment Methods 2014 :105-16. [Google Scholar]
[19]. Ward LD, Kellis M, HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and diseaseNucleic Acids Research 2016 44(D1):D877-D81. [Google Scholar]
[20]. Ding Y, Chan CY, Lawrence CE, RNA secondary structure prediction by centroids in a Boltzmann weighted ensembleRNA 2005 11(8):1157-66. [Google Scholar]
[21]. Shinoda G, Shyh-Chang N, Soysa T, Zhu H, Seligson MT, Shah SP, Fetal deficiency of Lin28 programs life-long aberrations in growth and glucose metabolismStem Cells 2013 31(8):1563-73. [Google Scholar]
[22]. McCauley JL, Kenealy SJ, Margulies EH, Schnetz-Boutaud N, Gregory SG, Hauser SL, SNPs in Multi-species Conserved Sequences (MCS) as useful markers in association studies: a practical approachBMC Genomics 2007 8(1):266 [Google Scholar]
[23]. Ridolfi E, Fenoglio C, Cantoni C, Calvi A, De Riz M, Pietroboni A, Expression and genetic analysis of microRNAs involved in multiple sclerosisInternational Journal of Molecular Sciences 2013 14(3):4375-84. [Google Scholar]
[24]. North B, Curtis D, Martin E, Lai E, Roses A, Sham P, Further investigation of linkage disequilibrium SNPs and their ability to identify associated susceptibility lociAnnals of Human Genetics 2004 68(3):240-48. [Google Scholar]